Abstract
New biomarkers are becoming available for the identification of patients who are suitable for the future discontinuation of ABL TKI treatment. We have shown a specific association between the BIM deletion polymorphism and the clinical outcome after ABL TKI treatment discontinuation in patients with long-lasting molecularly undetectable residual disease. Thirty-two CML patients were included in this study. The cumulative incidence of loss of MMR was estimated as 25.0% at 12 months and 31.3% at 24 months. We found that the BIM deletion polymorphism is a better predictor than BIM SNP of early molecular relapse after TKI treatment discontinuation in CML patients. We have reached the final stage of establishing the criteria for the discontinuation of ABL TKI treatment in CML patients. We are beginning to understand the strategies required for the successful discontinuation of ABL TKI treatment.
Key words
BIM, ABL TKIs, CML
Imatinib and the new ABL tyrosine kinase inhibitors (TKIs) have become the most successful class of targeted therapies for cancer, exceeding all projected survival expectations. With ABL TKI treatment, the annual all-cause mortality rate of chronic myeloid leukemia (CML) declined to 2% compared with a historical rate of 10% to 20%, and the estimated 10-year survival rate increased from 20% to greater than 80% [1]. The discontinuation of imatinib is currently being investigated for patients with CML who are in prolonged complete molecular response (CMR) [2,3]. In 2010, Mahon et al. reported a pilot study evaluating the feasibility and safety of imatinib discontinuation in patients with CMR of longer than 2 years and observed that half of the patients experienced molecular relapse during the 6 months after treatment discontinuation [2]. No late relapse was observed in the remaining patients during an extended follow-up period of longer than 4 years.Therefore, the discontinuation of ABL TKItreatmentmay indeed be possible in CML patients with prolonged CMR. However, no predictive prognostic factors for successful therapy discontinuation have yet been identified.
Recently, Ng et al. demonstrated that East Asian CML patients with a common deletion polymorphism in intron 2 of the BCL2L11 (BIM) gene had a lower response to TKIs than to those without the deletion [4]. Augis et al. also demonstrated that CML patients with the BIM SNP had a lower response to imatinib [5]. We therefore aimed to further our understandingof the predictive biomarkers for molecular relapse and non-relapse after ABL TKI treatment discontinuation. This study was approved by the institutional review board of Tokyo Medical University (no. 1655: approved on January 28, 2011). Patients in CMR receiving TKI treatment were eligible for inclusion in this study. Molecular relapse was defined as a loss of the major molecular response (MMR). Genomic DNA of the patients was obtained from whole blood using the EZ1 DNA Blood 350 l kit (Qiagen, Valencia, USA) and was subjected to polymerase chain reaction amplification using primers designed to detect a deletion site (2,903 bp) in intron 2 (deletion polymorphism) and a silent SNP in exon5 (c465C>T) [6].
Thirty-two CML patients (17 men and 15 women; median age: 57.5 years) were included in this study (Sokal category: low, 24 patients; intermediate, 7 patients; high, 1 patient). Six patients were treated with interferon-α (IFNα) before TKI treatment, and 3 were treated with IFNα after discontinuation of TKI treatment. Five patients (16%) had the BIM deletion polymorphism, and 7 patients (22%) had the BIM SNP. Furthermore, none of the patients had both BIM genetic variants. The median duration from TKI treatment initiation to discontinuation was 80.5 months (range: 22 to 138 months); median duration of CMR before TKI treatment discontinuation was 43.0 months (range: 5 to 97 months). Nine patients showed a loss of MMR, 8 patients relapsed within 6 months, and 1 patient showed a late relapse at 24 months after discontinuation. The cumulative incidence of loss of MMR was estimated as 25.0% at 12 months and 31.3% at 24 months (Figure 1A). The fluctuation of BCR-ABL transcript levels below the MMR threshold (>2 consecutive positive values) was observed in 3.13% of the patients at 24 months after ABL TKI treatment discontinuation. Treatment-free remission was estimated as 75.0% at 12 months and 68.8% at 24 months. The median period of restoration of the second CMR was 6.0 months in re-treated patients. None of the patients died during the follow-up period. TKI-free remission was estimated as 68.8% at 30 months (Figure 1A). A significant difference in the BIM deletion polymorphism was only observed between the patients who maintained MMR and those who lost MMR (p = 0.0031; long-rank test) (Figure 1B). No significant difference was observed in the BIM SNP (p = 0.0757; long-rank test) (Figure 1C), prior IFNα therapy, time to a complete cytogenetic response, time to MMR, and time to CMR between relapsing and non-relapsing patients.
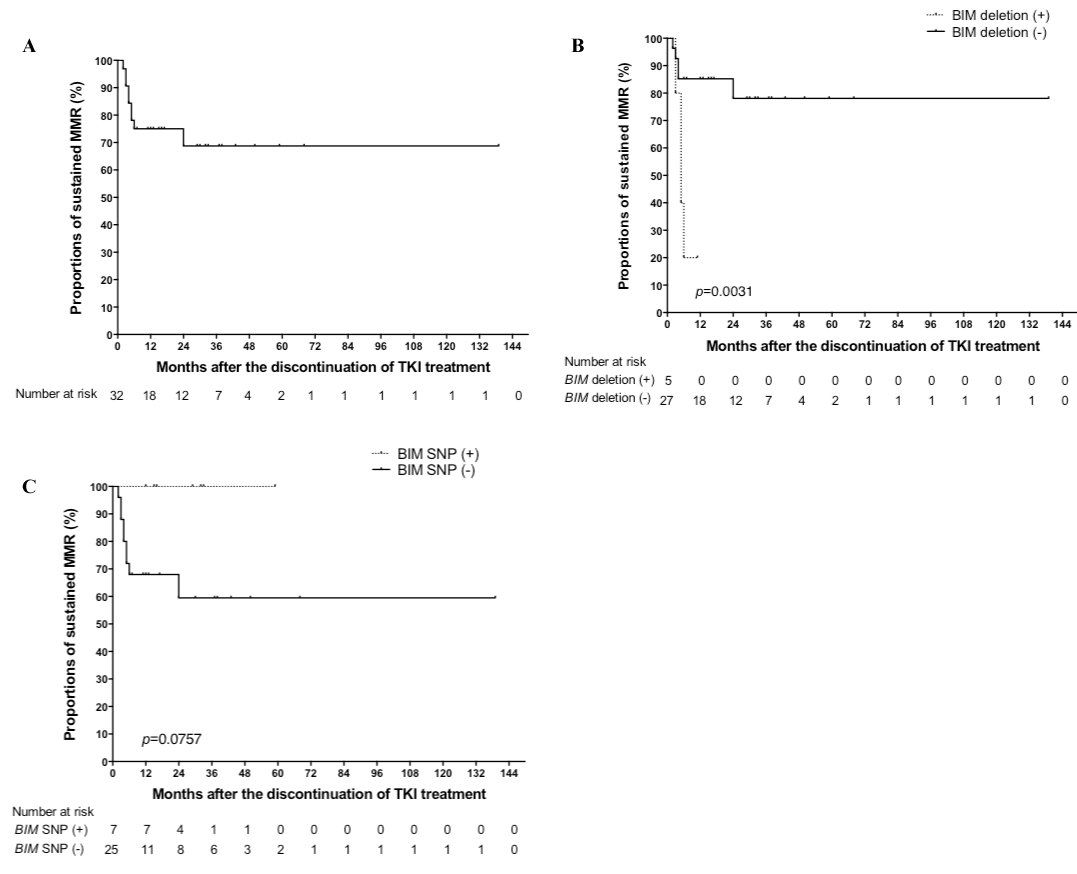
Figure 1. Kaplan-Meier estimates of major molecular remission after discontinuation of tyrosine kinase inhibitor treatment in all 32 patients (A), according to the BIMdeletion polymorphism (B), and according to the BIMSNP (C).
2021 Copyright OAT. All rights reserv
In this study, we found that the BIM deletion polymorphism is a better predictor than BIM SNP of early molecular relapse after TKI treatment discontinuation in CML patients. However, although the BIM deletion polymorphism is found specifically in East Asian patients [4], early molecular relapse was also observed in Caucasian patients. Genetic variations of other BH3-only proteins (for example, PUMA and NOXA) may alsobe different between patients of different races.
New biomarkers are becoming available for the identification of patients who are suitable for the future discontinuation of ABL TKI treatment. Here we have shown a specific association between the BIM deletion polymorphism and the clinical outcome after ABL TKI treatment discontinuation in patients with long-lasting molecularly undetectable residual disease. We have reached the final stage of establishing the criteria for the discontinuation of ABL TKI treatment in CML patients. We are also beginning to understand the strategies required for the successful discontinuation of ABL TKI treatment. Analysis of the BIM deletion polymorphism in CML patients is expected to be useful for predicting their early molecular relapse after the discontinuation of ABL TKI treatment.
References:
- Kantarjian H, O'Brien S (2012) The chronic myeloid leukenias. In: Goldman L, Schafer AI. Goldman's Cecil Medicine. (24thEdn). Philadelphia, PA Elsevier Saunders: 1209-1218.
- Mahon FX, Réa D, Guilhot J, Guilhot F, Huguet F, et al. (2010) Discontinuation of imatinib in patients with chronic myeloid leukaemia who have maintained complete molecular remission for at least 2 years: the prospective, multicentre Stop Imatinib (STIM) trial. Lancet Oncol 11: 1029-1035. [Crossref]
- Rousselot P, Charbonnier A, Cony-Makhoul P, Agape P, Nicolini FE, et al. (2014) Loss of major molecular response as a trigger for restarting tyrosine kinase inhibitor therapy in patients with chronic-phase chronic myelogenousleukemia who have stopped imatinib after durable undetectable disease. J Clin Oncol 32: 424-430. [Crossref]
- Ng KP, Hillmer AM, Chuah CT, Juan WC, Ko TK, et al. (2012) A common BIM deletion polymorphism mediates intrinsic resistance and inferior responses to tyrosine kinase inhibitors in cancer. Nat Med 18: 521-528. [Crossref]
- Augis V, Airiau K, Josselin M, Turcq B, Mahon FX, et al. (2013)A single nucleotide polymorphism in cBIM is associated with a slower achievement of major molecular response in chronic myeloid leukaemia treated with imatinib. PLoS One 8: e78582. [Crossref]
- Ohyashiki K, Tadokoro K, Yamaguchi T, Katagiri S, Umezu T, et al. (2014) Detection of BIM (BCL2L11) Polymorphic Variants in Chronic Myeloid Leukemia by Q-Invader Assay and Their Clinical Significance. J Hematol Transfus 2: 1032.

