Abstract
5-Fluorouracil (5-FU) is widely used in cancer therapy, either alone or in combination with other anti-cancer drugs. However, poor membrane permeability and a short half-life (5-20 min) due to rapid metabolism in the body necessitate the continuous administration of high doses of 5-FU to maintain the minimum therapeutic serum concentration. This is associated with significant side effects and a possibility of severe toxic effects. This study aimed to formulate 5-FU-loaded pH-sensitive liposomal nanoparticles (pHLNps-5-FU) and evaluate 5-FU release characteristics and anti-cancer effect of pHLNps-5-FU. Particle size and zeta potential were determined using a particle size analyzer. The release patterns of pHLNps-5-FU formulations were evaluated at 37°C at pH 3, 5, 6.5, and 7.4, while drug release kinetics of 5-FU from a pHLNp3_5-FU formulation were determined at pH 3 and 7.4 at different time points (37°C). Cell viability and clonogenic studies were conducted to evaluate the effectiveness of pHLNps-5-FU against HCT-116 and HT-29 cell lines while cellular uptake of rhodamine-labeled pHLNps-5-FU was determined by flow cytometry and confocal imaging. The average sizes of the pHLNp1_5-FU, pHLNp2_5-FU and pHLNp3_5-FU liposomes were 200 nm ± 9.8 nm, 181.9 nm ± 9.1 nm, and 164.3 nm ± 8.4 nm respectively. In vitro drug release of 5-FU from different pHLNps-5-FU formulations was the highest at pH 3.8. Both cell lines treated with pHLNps-5-FU exhibited reduced viability, two- or three-fold lower than that of 5-FU-treated cells. Flow cytometry and confocal imaging confirmed high uptake of rhodamine-labeled pHLNps-5-FU in both cell lines. The drug release profile of the chosen pHLNp3-5-FU formulation was optimal at pH 3 and had the poorest release profile at pH 7.4. The release profile of pHLNp3-5-FU showed that 5-FU release was two-fold higher at pH 3 than that at pH 7.4. This study demonstrates that pHLNp3-5-FU may be a potential candidate for the treatment of colorectal cancer.
Key words
pH-Sensitive liposomes, nanoparticles, 5-Fluorouracil (5-FU), colorectal cancer, clonogenic assay
Introduction
Colorectal cancer (CRC) is the formation of abnormal growths or polyps in tissues that line the colon or rectum. CRC is the third most commonly diagnosed cancer worldwide, and the second most common cause of cancer-related death in the western world [1-5]. It is estimated that 93,090 new cases of colon cancer and 39,610 of rectal cancer will be diagnosed in the year 2015; 49,700 cancer-related deaths are expected to be attributed to colorectal cancer [6].
Apart from radiation, surgery, and biologic therapies (immunotherapy and hormonal therapy), cytotoxic drugs comprise the majority of chemotherapy regimens used clinically for the treatment of cancers [7]. Most chemotherapeutics in current use interfere with cell replication in some manner, either by acting like nucleoside analogs (leading to S-phase arrest), or damaging deoxyribonucleic acid (DNA). As cancer cells undergo rapid cell divisions, they are generally more susceptible to these drugs than normal cells. In addition, cancer cells often lack the ability to recognize and/or repair DNA damage that leads to improper replication of cellular DNA and eventually causes cell death. The negative impact of this therapy is widespread, including chemotherapeutic drug resistance and interference with normal cell division, causing profound drug toxicity [7]. The next-generation targeted drugs may have few side effects since they are designed to target specific factors more accurately, such as the overexpressed receptors or proteins that are barely or never present in normal cells. These drugs are still largely in clinical trial and not available for most patients or they are barely in common clinical use, with a few well-known exceptions (i.e. human epidermal growth factor receptor 2 (HER2)-targeting trastuzumab and various epidermal growth factor receptor (EGFR) inhibitors [7,8].
5-FU is an antimetabolite of the pyrimidine analogue type with a broad spectrum of activity against solid tumors, either alone or in combination with other chemotherapy regimens. Due to its structure, which is a base analogue that mimics both uracil and thymine, 5-FU interferes with nucleoside metabolism by incorporating into ribonucleic acid (RNA) and DNA, leading to cytotoxicity and cell death. Despite its therapeutic efficacy, 5-FU has limitations that include: i) tumor cell resistance; for example, overall response rate of advanced CRC to 5-FU alone is 10-20% while that of 5-FU in combination with other antitumor drugs is 40-45% [9], and ii) short biological half-life (5 to 20 min), which is owing to rapid metabolism in the body; therefore, the maintenance of therapeutic serum concentration often requires continuous administration of high doses, which may lead to severe toxicity [7,10].
These issues can be mitigated by formulating 5-FU in a delivery system that causes accumulation of the drug in tumor regions and increases exposure time in cancer cells. A suitable 5-FU delivery system with these characteristics should have the following properties: a) physical stability; b) small size to allow capillary distribution and uniform perfusion at the desired target site; c) the ability to carry adequate amount of the drug with negligible or low drug leakage, d) the ability to protect 5-FU from degradation, and e) controllable (or predictable) 5-FU release rates from the carrier at the desired target site [11,12].
Recently, the focus of liposomal research has been the development of strategies to increase the ability of liposomes to mediate intracellular delivery of biologically active molecules [13]. This has led to the emergence of liposomes called stealth liposomes (liposomes sterically stabilized with polyethylene glycol (PEG)). Stealth liposomes are more suitable than polymers as a delivery system for 5-FU because they are stable, biocompatible, biodegradable, lack immunogenic response, and overall possess the properties of a good delivery system already described above, while polymers may cause serious toxicity with innate breakdown products. In addition, the Food and Drug Administration (FDA) has previously approved stealth liposomes for the delivery of doxorubicin for the treatment of breast cancer and ovarian cancer [14].
pH-sensitive liposomes are a modified form of stealth liposomes that are stable at physiological pH (pH 7.4) but undergo destabilization under acidic conditions. These are reported to be more efficient in delivering anti-cancer drugs than conventional or long-circulating liposomes owing to their fusogenic property [13]. pH-sensitive stealth liposomes functionalized with an appropriate moiety (i.e. an antibody) targeted to receptor expressing cancer cells such as EGFR have been shown to significantly increase the intracellular delivery of their liposomal content [15].
In this study, we describe the development of 5-FU-loaded pH-sensitive liposomal nanoparticles with surface-modified anti-EGFR antibody-conjugated pHLNps-5-FU, and provide an in vitro evaluation of its therapeutic potential for cancer chemotherapy. The in vitro uptake and the cytotoxic activity of pHLNps-5-FU have been analyzed and compared with 5-FU by using HCT-116 and HT-29 cell lines, which are established cell culture models of CRC cancer.
Materials and methods
Materials
All chemicals including 5-FU and reagents were purchased from Sigma-Aldrich (St. Louis, Missouri, USA). Dioleoylphosphatidylethanolamine (DOPE), cholesterylhemisuccinate (CHEMS), phosphatidylcholine (L-α-PC), 1,2-dioleoyl-3-trimethylammoniumpropane (DOTAP), cholesterol (CH), 1,2-distearoyl-sn-glycero-3- phosphoethanolamine-N-[amino(polyethyleneglycol)-2000] (DSPE-PEG), and 1,2-distearoyl-sn-glycero-3-phosphoethanolamine-N-[folate(polyethyleneglycol)-2000](DSPE-PEG-FA) lipids were all obtained from Avanti Polar Lipids, Inc. (Alabaster, AL, USA). Cancer cell lines, HCT-116 and HT-29, were obtained from American Type Culture Collection (ATCC) in January, 2013.
Preparation of 5-FU loaded pH-sensitive liposomal nanoparticles
Different pH-sensitive liposomal nanoparticles containing 5-FU and pHLNp (pHLNp1-5-FU, pHLNp2-5-FU, and pHLNp3-5-FU) were prepared by thin film hydration method [16]. Briefly, varying amounts of lipids were measured in different molar ratios (Table 1) and placed in different round-bottom flasks. The lipids were then dissolved in chloroform and mixed thoroughly. The chloroform was then removed by passing nitrogen gas through the inner side of a flask in a fume hood. The thin film obtained was further dried under vacuum overnight to remove any residuals. The dried film was then hydrated at a temperature above the transition temperature of the lipid (60°C) with 2 ml of phosphate buffer solution (PBS) pH 7.4 containing 19 µM 5-FU. The hydrated film was then vortexed for 1 min and bath sonicated for 5 min. The resulting multi-laminar liposomal vesicles were then extruded through a 200-nm polycarbonate filter membrane to further reduce the size. The free 5-FU was finally removed by dialysis against PBS for 24 hr using a 12 kDa molecular weight cutoff dialysis tube.
Table 1. Characterization of 5-FU loaded-pH-sensitive liposomal nanoparticles
|
Formulation
|
Lipid composition
|
Molar ratio
|
Mean particle Size (nm)
|
Zeta potential (mV)
|
Entrapment efficiency (%)
|
|
pHLNP1
pHLNP1-5-FU
|
DOPE:CHEM:DSPE-PEG2000
DOPE:CHEM:DSPE-PEG2000
|
50:30:20
50:30:20
|
189.3 ± 7.7
200.0 ± 9.8
|
2.34 ± 1.5
5.16 ± 1.4
|
-
4.74
|
|
pHLNP2
pHLNP2-5-FU
|
PC:CHEM:TWEEN 80:DSPE-PEG2000
PC:CHEM:TWEEN 80:DSPE-PEG2000
|
40:40:10:10
40:40:10:10
|
155.4 ± 8.5
181.9 ± 9.1
|
0.59 ± 1.3
1.98 ± 0.5
|
-
3.25
|
|
pHLNP3
pHLNP3-5-FU
|
CHEM:CH:TWEEN 20:DSPE-PEG2000
CHEM:CH:TWEEN 20:DSPE-PEG2000
|
60:20:10:10
60:20:10:10
|
136.1 ± 10.2
164.0 ± 8.4
|
1.30 ± 0.8
1.23 ± 0.8
|
-
54.17
|
Data expressed as mean ± SEM, n = 3 (Dioleoylphosphatidylethanolamine (DOPE), cholesterylhemisuccinate (CHEMS), cholesterol (CH), 1,2-distearoyl-sn-glycero-3- phosphoethanolamine-N-[amino(polyethyleneglycol)-2000] (DSPE-PEG2000),phosphatidylcholine (PC))
Characterization of pH-sensitive liposomal nanoparticles
Size measurement: The particle size and zeta potential of the blanks, pHLNp1, pHLNp2, and pHLNp3 and their corresponding loaded 5-FU, pHLNp1-5-FU, pHLNp2-5-FU, and pHLNp3-5-FU were determined by dynamic light scattering using a zeta potential/Particle Sizer Instrument (NICOMP™ 380 ZLS) (Table 1). All measurements were performed in triplicates and the results were reported in mean diameter ± SEM.
Entrapment efficiency (EE %): Ten milligrams of lyophilized pHLNp1-5-FU, pHLNp2-5-FU, or pHLNp3-5-FU was suspended in 2 ml of PBS (pH 7.4). The liposomal suspension was disrupted by adding 100 µL of 30% Triton X-100, gently mixed for 2 min, and centrifuged at 6,000 rpm at room temperature for 5 min. The supernatant was collected and analyzed for 5-FU using reverse phase high-performance liquid chromatography (HPLC). The reverse phase HPLC: the mobile phase solution consisting of 95% PBS and 5% of methanol was prepared and filtered according to method described [17]. The internal standard or sample injection volume was 20 µL, which was pumped through a XB-C18 column at a flow rate of 1.0 mL/min (250 mm × 4.6 mm; Agilent, Santa Clara, CA) at room temperature. 5-FU was detected at 270 nm with Waters 996 photodiode array detector (Waters, Columbia Maryland. USA). The entrapment efficiency was calculated according to the following equation:
(1)
FTIR analysis: Based on HPLC analysis, the pHLNp3 formulation was found to have the highest EE (%) of 5-FU (Table 1). FTIR analysis was further conducted on the pHLNp3 to confirm the incorporation of 5-FU. Physical mixture (lyophilized blank pHLNp3 and 5-FU), lyophilized blank pHLNp3, 5-FU, and lyophilized pHLNP3-5-FU were analyzed in spectra range of 740–4000 cm-1 using FTIR spectrophotometer (PerkinElmer Life and Analytical Sciences, Connecticut, USA). The acquired spectra were then used to determine the presence of 5-FU in pHLNP3.
Evaluation of in-vitro drug release
5-FU release at different pH values: Buffer solutions of different pH values (3.0, 5.0, 6.5 and 7.4) were prepared and 1 ml of liposomal nanoparticle suspension containing 2.5 mg/ml of pHLNp1-5-FU, pHLNp2-5-FU, or pHLNp3-5-FU was placed in a dialysis bag; this was immersed completely in different pH solutions and stirred continuously (100 rpm) for 24 hr at 37°C. After 24 hr, 500 µL of solution was removed from each receiver chamber and analyzed for the presence of 5-FU using reverse phase HPLC as described above.
5-FU release at different time points: A buffer solution of a pH 3.0 was prepared and 1 ml of liposomal nanoparticle suspensions containing 2.5 mg/ml of pHLNp1-5-FU, pHLNp2-5-FU, or pHLNp3-5-FU was placed in dialysis bag and immersed completely in solution of pH 3 and stirred at 100 rpm continuously for 24 hr at 37°C. At predetermined time intervals of 1, 5, 10, 15, 30, 60, 120, 240, 480, 720, and 1440 min, 500 µL of receiver solution was sampled out and replaced with equal volume of fresh PBS at 37°C. Amount of 5-FU present in each sampled solution was determined by reverse phase HPLC.
Cell viability
The in vitro cytotoxicity of free 5-FU, pHLNp1-5-FU, pHLNp2-5-FU, or pHLNp3-5-FU was evaluated using HCT-116 and HT-29 colon cancer cell lines. The HCT-116 and HT-29 cell lines were seeded into 12-well plates at a density of 5 x 103 cells per well and cultured in DMEM/F12 media supplemented with 2 mM L-glutamine, 10 mM HEPES, 10% fetal bovine serum (FBS), and 1% penicillin/streptomycin. The cells were treated with different concentrations of 5-FU or its equivalent in pHLNp1-5-FU, pHLNp2-5-FU or pHLNp3-5-FU after they had reached 75% confluence. After 48 hr, the experiments were terminated and the cells were detached, stained with 2% trypan blue, and counted with an automated cell counter (Bio-Rad TC- 20™). The cell viability (%) relative to the control was determined.
Cellular uptake
Confocal imaging: HCT-116 and HT-29 cancer cells were grown in 6-well plates (with cover slips) at a cell density of 2 x 103, for 24 hr at 37°C. The cells were then treated with rhodamine-labeled pHLNPs in growth media (Rho-pHLNp1, Rho-pHLNp2, or Rho-pHLNp3). After 3 hr, Rho-pHLNp1, Rho-pHLNp2, or Rho-pHLNp3 was removed and the cells gently washed twice with PBS (pH 7.4). Next 0.75 µg/ml of 4,6-diamidino-2-phenylindole (DAPI) was added for nuclear staining; finally, the cells were fixed, using 4% paraformaldehyde, then mounted and imaged using Leica SP2 Multiphoton system.
Lysosomal delivery of Lyso Tracker Red DND-99 by pHLNP: HT-29 and HCT-116 cells were seeded on cover slips at a density of 2.5 x 105 per well in a 6-well plate. After 24 hr cultivation, cells were incubated with 500 µg/ml of Lucifer yellow-labeled pHLNP3-5-FU (LY-pHLNP3-5-FU) for 4 h at 37°C, and then incubated with LysoTracker Red DND-99 (200 nM) for 1 hr. The cells were then washed three times with cold PBS (pH 7.4), fixed using 4% paraformaldehyde, and the cover slips were mounted cell-side down with slides and viewed using a Zeiss LSM 880 Confocal Microscope [18].
Flow cytometry: To determine 5-FU-loaded liposomal nanoparticle uptake by the cells, HCT-116 and HT-29 cells were plated onto 6-well plates with a density of 5 × 105 and cultured in growth media until 75% confluence. Cells were incubated with the different rhodamine-labeled liposomes (Rho-pHLNp1, Rho-pHLNp2 or Rho-pHLNp3) for 24 hr at 37°C. After incubation, the cells were detached from the culture plate with 0.25% trypsin-EDTA solution, washed three times with PBS, and centrifuged at 3,000 rpm for 5 min. Finally, the cells were re-suspended in 500 µl PBS, fixed with 4% paraformaldehyde, and kept on ice until analysis using a BD FACSCanto™ Analyzer and a BD FACSAria™ Cell Sorter (BD Biosciences)
Colony formation assay
For colony assay, HCT-116 and HT-29 cell lines were seeded into T-25cm2 culture flask at a density of 5 × 105 cells and cultured in DMEM/F12 medium supplemented with 2 mM L-glutamine, 10 mM HEPES, 10% FBS, and 1% penicillin/streptomycin. After the cells reached 75% confluency, they were exposed to different concentrations of free 5-FU and pHLNP3-5-FU. After 48 hr exposure and two treatments, the experiment was terminated, cells harvested, and then re-plated onto 6-well plates at a density of 200, 500, and 1,000 cells per well, and incubated with growth medium. After the control cells reached 75% confluence, the experiment was terminated by fixing and staining the plates with 0.5% crystal violet solution. The stained colonies (fifty per colony) were counted using a Jenco™ Stereomicroscope; plating efficiency (PE) and surviving fraction (SF) were calculated, and a graph of survival curve graph was generated [19].
Statistical analysis
All experiments were performed in triplicates and analyzed using GraphPad Prism software (GraphPad Software Inc., La Jolla, CA. USA). The differences between the 5-FU treatment group and that of pHLNp1-5-FU, pHLNp2-5-FU or pHLNp3-5-FU were determined using Student’s paired t-test and considered significant at p<0.05.
Results
Characterization of 5-FU pH-sensitive liposomal nanoparticles
Mean particle sizes of blank liposomal nanoparticles pHLNp1, pHLNp2, and pHLNp3 were 189.3 nm ± 7.7 nm, 155.4 nm ± 8.5 nm, and 136 nm ± 10.2 nm, respectively, while the mean sizes of 5-FU-loaded liposomal nanoparticles; pHLNp1_5-FU, pHLNp2_5-FU and pHLNp3_5-FU were found to be 200 nm ± 9.8 nm, 181.9 nm ± 9.1 nm, and 164.3 nm ± 8.4 nm, respectively (Table 1). Of all the formulations, pHLNp3_5-FU had the highest 5-FU entrapment efficiency (EE) of 54.17%, whereas pHLNp1_5-FU and pHLNp2_5-FU showed comparable 5-FU EE of 3.25 and 4.74%, respectively. The zeta potential values of pHLNp1_5-FU, pHLNp2_5-FU, and pHLNp3_5-FU were 5.16 ± 1.4, 1.98 ± 0.5, and 1.23 ± 0.8, respectively (Table 1).
FTIR Analysis
Although FTIR analysis is not confirmatory approach to fully determine entrapment of 5-FU in the liposomal carrier, it could be used to assess the association of 5-FU and the carrier. A careful examination of 5-FU spectrum revealed completely different absorption peaks when compared with blank or empty pHLNPs carrier (Figure 1 A and D). Further, neither 5-FU (Figure A) nor blank pHLNPs (Figure 1 D) absorption peaks were not similar to that of pHLNPs-5-FU (Figure 1 B) peaks. The unique features of pHLNPs-5-FU peaks clearly showed a close association between 5-FU and pHLNPs carriers. The FTIR spectra of pure 5-FU showed a –N-H stretch at 3120 cm-1 and –C=O stretch at 1644.9 cm-1. These same stretches were also seen in the pHLNPs-5-FU sample that was analyzed, therefore confirming the interaction of 5-FU and the liposomal nanocarrier.
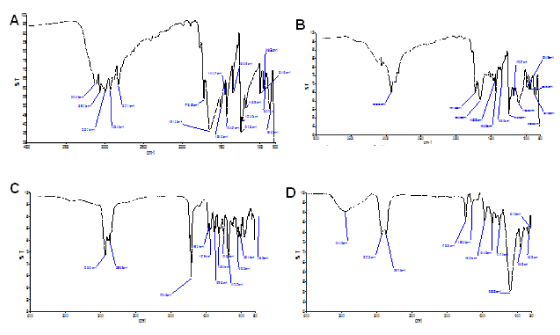
Figure 1. FTIR Spectra.
A; 5-FU. B; pHLNps-5-FU. C; Physical Mixture (CHEM, CH, TWEEN 20, DSPE-PEG2000). D; Blank pHLNPs.
In vitro 5-FU release
5-FU release at different pH values
The percent of 5-FU released decreased as pH increased from 3 to 7.4. This was a general trend observed in all of pHLNPs-5-FU formulations as shown in Figure 2A. However, pHLNP3-5-FU formulation had a much greater release of 5-FU (Figure 2B) compared to pHLNP1-5-FU and pHLNP2-5-FU (Figure 2A). Among all the formulations, pHLNP3-5-FU had the highest release of 5-FU (30%) at pH 3, compared to 5-FU release at pH 7.4 (15%). Further, pHLNP3-5-FU was the most pH-sensitive formulation compared with pHLNP1-5-FU and pHLNP2-5-FU at pH 3.
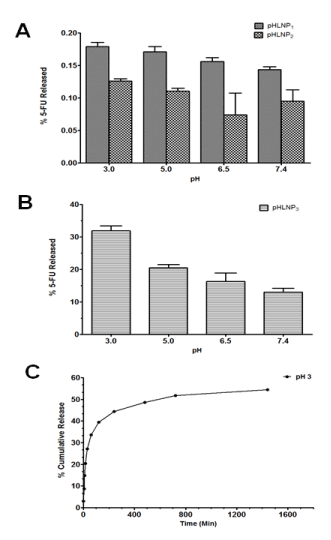
Figure 2. In-vitro release of 5-FU.
A and B; in-vitro release of 5-FU from pHLNP1, pHLNP2 and pHLNP3 at different pH values. C; Cumulative 5-FU release from pHLNP3 at pH 3.
5-FU release at different time points
The cumulative release profile of 5-FU from pHLNP3-5-FU at pH 3 was biphasic, as shown in Figure 3C. There was an initial rapid release of 35% of 5-FU for the first 100 min, followed by the release of an additional 15% over a period of 700 min (from 100 to 800 min). After 800 min, 5-FU release did not significantly increase with increasing time; only 3% of 5-FU was released from 800 min to 1,440 min (Figure 2C).
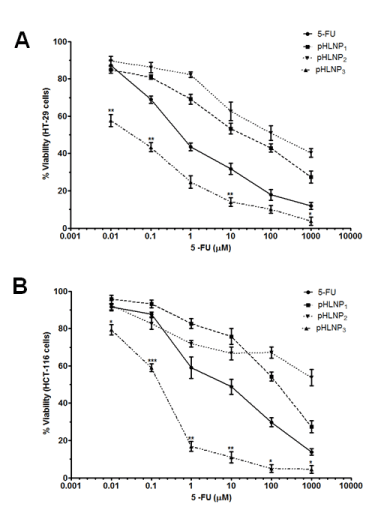
Figure 3. Effects of 5-FU and pHLNPs-5-FU on cell viability
Cytotoxicity of 5-FU and pHLNP1-5-FU, pHLNP2-5-FU and pHLNP3-5-FU on) HT-29 (A) and HCT-116 (B) cell lines. (5-FU vs. pHLNP3-5-FU; *p<0.05, **p<0.01, ***p<0.001). P-value was calculated by Student’s t-test. Data represents mean ± SD, n=3.
Cell viability
Figure 3 shows the effect of increasing concentration of 5-FU, pHLNP1-5-FU, pHLNP2-5-FU, and pHLNP3-5-FU on the viability of HCT-116 and HT-29 cancer cell lines. As shown in Figure 3A, pHLNP3-5-FU significantly inhibited HT-29 cell growth at all concentrations compared to free 5-FU. Further, pHLNP3-5-FU was two- to three-fold more effective than pHLNP1-5-FU or pHLNP2-5-FU. As shown in Figure 3B, pHLNP3-5-FU was the most effective formulation against HCT-116 cell lines, as compared to free 5-FU, pHLNP1-5-FU, and pHLNP2-5-FU.
2021 Copyright OAT. All rights reserv
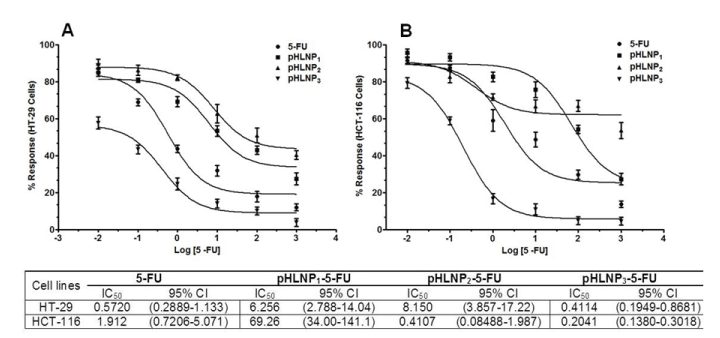
Figure 4. Non-linear curve fitting for dose response curve
Dose response of 5-FU and pHLNP1-5-FU, pHLNP2-5-FU and pHLNP3-5-FU on HT-29 (A) and HCT-116 (B) cell lines and their corresponding IC50 values (µM).
For example, 1 µM 5-FU-loaded pHLNP3 was four-fold more effective in inhibiting HCT-116 cell growth than pHLNP1-5-FU or pHLNP2-5-FU, and also three-fold more effective than free 5-FU in HCT-116 cell growth inhibition. To measure the effectiveness of free 5-FU and the 3 pHLNPs-5-FU formulations in inhibiting both HT-29 and HCT-116 cells’ growth, we modeled the data obtained from cell viability testing using varying concentrations of pHLNP1-5-FU, pHLNP2-5-FU, and pHLNP3-5-FU to generate half-maximal inhibitory concentrations (IC50) as shown in Figure 4. As expected, pHLNP3-5-FU was the most effective formulation, with IC50 values of 0.4114 (µM) on HT-29 cells and 0.2041(µM) on HCT-116 cells. Conversely, pHLNP1-5-FU (IC50 = 6.256 (µM) against HT-29 cells and IC50 = 69.26 µM against HCT-116 cells) was found to be the most ineffective formulation to inhibit HCT-116 cells growth while pHLNP2-5-FU (IC50 = 8.150 µM against HT-29 cells and IC50 = 0.4107 µM against HT-116 cells) was found to be least effective formulation against HT-29 cells.
Cellular uptake
Confocal imaging
Cellular uptake of pHLNP formulations was determined by treating both HT-29 and HCT-116 cells for 3 hr at 37°C with DOPE-Rho-conjugated pHLNPs; Rho-pHLNP1, Rho-pHLNP2, and Rho-pHLNP3. Confocal images of HT-29 and HCT-116 cells showed significant uptake of Rho-pHLNP1, Rho-pHLNP2, and Rho-pHLNP3 as shown in Figure 5. The merged images clearly showed that majority of internalized nanoparticles were localized in the cell nuclei.
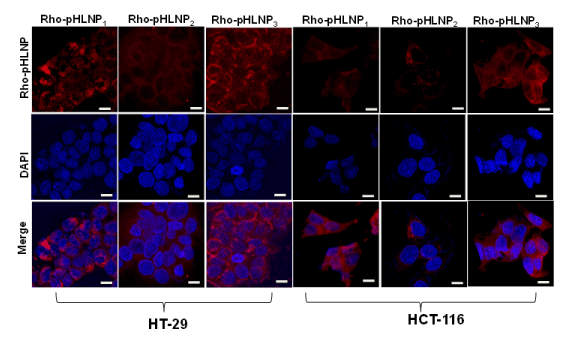
Figure 5. Confocal laser imaging
Cellular uptake of Rho-pHLNP1, Rho-pHLNP2 and Rho-pHLNP3 by HT-29 and HCT-116 cells after incubation for 3 hour. Nuclei were stained with DAPI (Scale bar = 20 µM).
Lysosomal delivery of Red DND-99 by pHLNP
We also investigated the internalization and intracellular fate of and counterstaining with LysoTracker Red DND-99. Figure 6A & D show that most of LY-pHLNP3-5-FU was taken up by cells (green color). LysoTracker Red DN-99 uptake by the cells was observed as red color (Figure 6B & E). The co-localization of LY-pHLNP3 and LysoTracker Red (Figure 6C & F) was followed and the resulting yellow color shows the accumulation of LY-pHLNP3-5-FU in the lysosomes. This was performed for both HT-29 and HCT-116 cell lines.
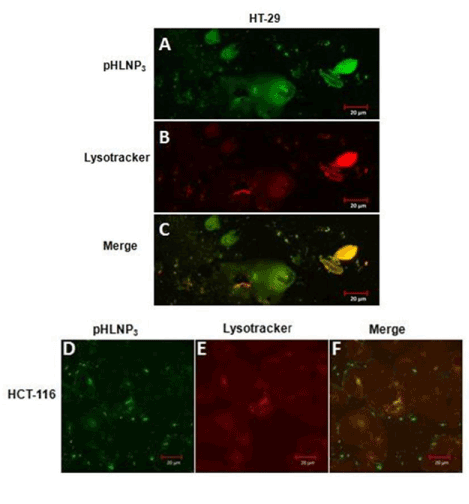
Figure 6. Endo-lysomal uptake of pHLNP3-5-FU in HT-29 and HCT-116 cells
For HT-29; (A) Internalization of pHLNP3-5-FU, (B) cell stained with lysotracker, (C) colocalization of internalized pHLNP3-5-FU and endo-lysosomes. For HCT-116; (D) Internalization of pHLNP3-5-FU, (E) cell stained with lysotracker, and (F) colocalization of internalized pHLNP3-5-FU and endo-lysosomes.
Flow cytometry
To further confirm the internalization of the formulations by HT-29 and HCT-116 cells, flow cytometry analysis was performed. Figure 7 shows cellular uptake of Rho-pHLNP1, Rho-pHLNP2, and Rho-pHLNP3 after the cells were incubated for 24 hours at 37°C. The results show significant uptake of the formulations by both cells, however HCT-116 cells exhibited greater cellular uptake of the formulations compared to HT-29 cells.
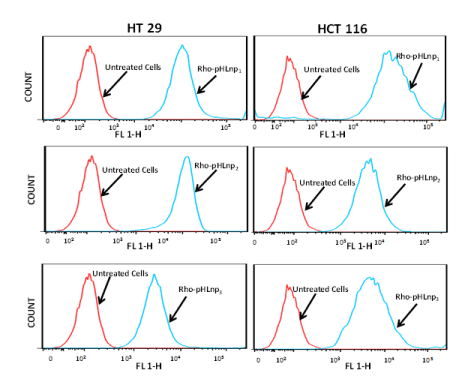
Figure 7. Flow cytometry analysis
Cellular uptake of Rho-pHLNP1, Rho-pHLNP2 and Rho-pHLNP3 by HT-29 and HCT-116 by flow cytometry analysis.
Colony assay
The proliferative properties of both HT-29 and HCT-116 cell lines were assessed after treatment with free 5-FU and pHLNp3_5-FU via a clonogenic assay method. Figure 8A shows a difference in percent survival of free 5-FU or pHLNP3-5-FU-treated HT-29 cells, as the concentrations of both free 5-FU and pHLNP3-5-FU were increased. As concentration increased from 0.01 to 10 µM, pHLNP3-5-FU was able to disrupt the proliferative property of the cancer cell better than free 5-FU. This is further confirmed by the survival curve shown in Figure 8B. Figure 9 shows a reduction of colony formation in HCT-116 cells as the doses of free 5-FU and pHLNP3-5-FU were increased. Figure 9B shows a survival curve with a similar pattern to that in Figure 8A. It also shows that pHLNP3-5-FU was more effective and significantly decreased the percent survival compared to free 5-FU at a concentration range of 0.01-10 µM.
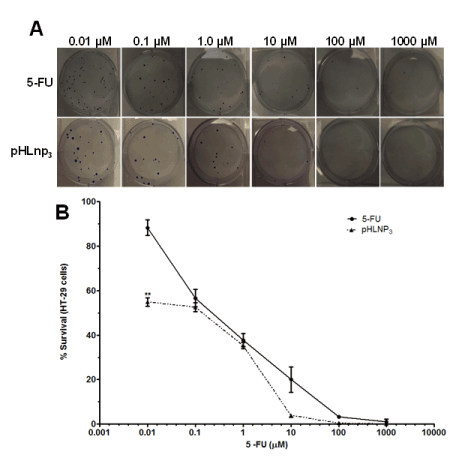
Figure 8. Clonogenic assay for HT-29 cells treated with 5-FU and pHLNP3-5-FU
Colony images (A) and survival curve (B) of after treatment with 5-FU and pHLNP3-5-FU. (5-FU vrs pHLNP3-5-FU; **p<0.01). P-value was calculated by Student’s t-test. Data represents mean ± SD, n=3
Put together, the results show that pHLNP3 could increase the delivery and anti-cancer activity of 5-FU in HT-29 and HCT-116 colorectal cancer cells.
Discussion
5-FU is a low molecular weight drug that acts as a thymidylate synthase inhibitor to block the synthesis of the pyrimidine thymidine, which is essential for DNA replication. It is currently used clinically to treat colorectal cancer either alone or in combination with other anticancer drugs [20]. For optimal therapeutic activity, 5-FU is administered continuously for an extended period of time to cancer patients due to a short half-life. However, due to lack of specificity, long-term therapy with 5-FU may lead to severe cardiotoxicity [21]. The aim of this study was to develop and study pH-sensitive liposomal nanoparticles loaded with 5-FU (pHLNps-5-FU), with surface-modified anti-EGFR antibody for the following purposes: i) increased 5-FU plasma circulation half-life, ii) increased anti-cancer activity of 5-FU, iii) reduced associated toxicity, and iv) improved specificity. A pH-sensitive liposome is an attractive delivery system due to the fact that the tumor site is relatively acidic compared to the normal tissue site, and the liposome can undergo destabilization to release its aqueous content under acidic condition [13,22].
Three pHLNps-5-FU formulations (pHLNP1-5-FU, pHLNP2-5-FU, and pHLNP3-5-FU) were prepared and characterized by size, zeta potential, and entrapment efficiency; pHLNP3-5-FU was chosen for further study because it had the highest entrapment efficiency (54.17%) (Table 1). This was attributed to the presence of a -CH moiety in the pHLNP3 formulation, which was necessary to prevent the leakage of encapsulated 5-FU from pHLNP3. CH molecules fill in a free space formed due to a kink in the chain of unsaturated lipids that was present in the liposome formulation [23]. In addition, CHEMS, a component of pHLNP3, is reported to cause destabilization of the liposomal membrane at acidic pH, which would enhance the deposition of 5-FU at the tumor site [24]. Entrapment of 5-FU in liposomal nanoparticles was also confirmed by FTIR. The FTIR spectrum of pure 5-FU showed a –N-H stretch at 3120 cm-1 and –C=O stretch at 1644.9 cm-1 (Figure 1). Similar stretches or absorption peaks were also observed in the pHLNPs3-5-FU spectrum suggesting the entrapment or presence of 5-FU in pHLNP3. As expected, in vitro release of 5-FU was highest in pHLNP3 compared to pHLNP1 and pHLNP2. This may be due to the fact that pHLNP3 has a much higher amount of CHEM lipid compared to pHLNP1 and pHLNP2, which is speculated to cause destabilization of the formulation and improve release of 5-FU under acidic conditions. The rapid disruption of pHLNP3 and release of 5-FU in the first 100 min at pH of 3 compared to pHLNP2 or pHLNP2 is attributed to the same reason [25].
The cytotoxicity of 5-FU was compared to pHLNP1-5-FU, pHLNP2-5-FU and pHLNP3-5-FU formulations on HT-29 and HCT-116 cancer cell lines to assess the most effective anti-cancer agent. Based on the viability and clonogenic results, HT-29 and HCT-116 cells were most sensitive to pHLNP3-5-FU compared to all other formulations, while pHLNP1-5-FU and pHLNP2-5-FU were less effective. The low anti-cancer activity of pHLNP1-5-FU and pHLNP2-5-FU is largely attributed to the low entrapment of 5-FU. Conversely, the high payload and increased delivery of 5-FU by pHLNP3, coupled with quick disruption of pHLNP3 under acidic conditions may significantly account for the enhanced anti-cancer activity of pHLNP3-5-FU. This was further supported by the IC50 results for pHLNP3-5-FU, which was the most effective agent against both HT-29 and HCT-116 cells. It was unclear why the IC50 for pHLNP2-5-FU was far better than that of 5-FU on HCT-116 cells.
Findings of flow cytometry and confocal studies revealed that the treatment of cells with the Rho-pHLNP1, Rho-pHLNP2, and Rho-pHLNP3 led to a comparable level of cell total Rho fluorescence intensity, which clearly indicates that the uptake of pHLNP1, pHLNP2, and pHLNP3 by HT-29 or HCT-116 cells was similar, although HCT-116 uptake of liposomal nanoparticles was slightly higher than that of HT-29 cells. While confocal and flow cytometry studies were conducted to assess the uptake of our formulations by the cells, delivery of pHLNP3 to the lysosomal compartment was studied to assess accumulation in the lysosomes. LY-pHLNP3-5-FU was chosen because of a high entrapment efficiency and extremely low IC50 values. The presence of LY-pHLNP3-5-FU in HT-29 or HCT-116 cells was observed by color green, while the lysosome compartment was stained the color red. To determine the accumulation of LY-pHLNP3-5-FU in the lysosome, the two images was merged and the yellow color (combination of green and red colors) was observed (Figure 6), confirming the presence of LY-pHLNP3-5-FU nanoparticles in the lysosome compartment.
In tumors, clonogenic assay measures the ability of individual cells to proliferate to form colonies of at least 50 or more cells, which is a critical metric of cell viability. Only clonogenic cells have the ability to cause recurrence or create metastasis [26]. Based on this, HT-29 and HCT-116 cells were tested for their ability to proliferate after treatment with 5-FU and pHLNP3-5-FU. The pHLNP1-5-FU and pHLNP2-5-FU formulations were not tested due to low 5-FU entrapment and poor inhibition capabilities made them poor candidates. The data suggest that pHLNP3-5-FU was more effective in rendering the cells incapable of proliferation compared to 5-FU (Figure 8 and 9). However, in comparing HT-29 and HCT-116 cells, it was evident that pHLNP3-5-FU had a much more pronounced effect in disrupting the proliferative property of HCT-116 than that of HT-29.

Figure 8. Clonogenic assay for HT-29 cells treated with 5-FU and pHLNP3-5-FU
Colony images (A) and survival curve (B) of after treatment with 5-FU and pHLNP3-5-FU. (5-FU vrs pHLNP3-5-FU; **p<0.01). P-value was calculated by Student’s t-test. Data represents mean ± SD, n=3
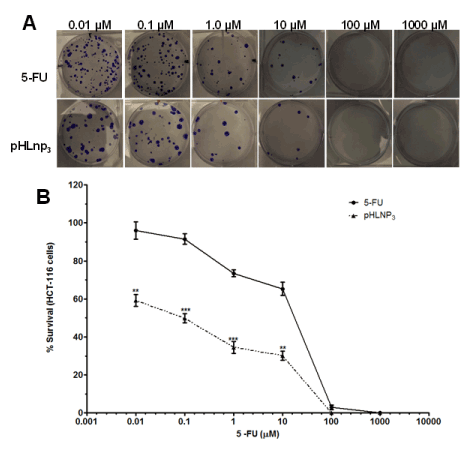
Figure 9. Clonogenic assay for HCT-116 cells treated with 5-FU and pHLNP3-5-FU
Colony images (A) and survival curve (B) of HCT-116 cancer cell lines after treatment with 5-FU and pHLNP3-5-FU. (5-FU vrs pHLNP3-5-FU; **p<0.01, ***p<0.001). P-value was calculated by Student’s t-test. Data represents mean ± SD, n=3
Conclusion
We have successfully formulated pH-sensitive thermo-sensitive liposomal nanoparticles, pHLNp3, which were very responsive to a pH below 4, exhibiting enhanced release of 5-FU under this condition. The pHLNp3-5-FU nanoparticles exhibited a stronger anti-cancer effect compared to 5-FU against HT-29 and HCT-116 cancer cells. The findings provide strong may evidence in support of a possible therapeutic application of pHLNp3 as a drug delivery system for 5-FU, which can overcome some of the limitations that 5-FU currently such as poor membrane permeability and short half-life. Currently, pHLNp3-5-FU is being studied in an animal model to evaluate in vivo efficacy on tumor growth.
Acknowledgement
Research reported in this publication was supported by the National Institute on Minority Health and Health Disparities (NIMHD) of the National Institutes of Health (NIH) under award numbers G12MD007582-28 and U54MD008149.
References
- Labianca R, Beretta GD, Kildani B, Milesi L, Merlin F, et al. (2010) Colon cancer. Crit Rev Oncol Hematol 74: 106-133. [Crossref]
- Foubert F, Matysiak-Budnik T, Touchefeu Y (2014) Options for metastatic colorectal cancer beyond the second line of treatment. Dig Liver Dis 46: 105-112. [Crossref]
- Gomez D, De Rosa A, Addison A, Brooks A, Malik HZ, et al. (2013) Cetuximab therapy in the treatment of metastatic colorectal cancer: the future frontier? Int J Surg 11: 507-513. [Crossref]
- Mishra J, Drummond J, Quazi SH, Karanki SS, Shaw JJ, et al. (2013) Prospective of colon cancer treatments and scope for combinatorial approach to enhanced cancer cell apoptosis. Crit Rev Oncol Hematol 86: 232-250. [Crossref]
- Longley DB, Allen WL, Johnston PG (2006) Drug resistance, predictive markers and pharmacogenomics in colorectal cancer. Biochim Biophys Acta 1766: 184-196. [Crossref]
- American Cancer Society: Cancer Facts & Figures 2015. 2015.
- Arias JL (2008) Novel strategies to improve the anticancer action of 5-fluorouracil by using drug delivery systems. Molecules 13: 2340-2369. [Crossref]
- Wong HL, Bendayan R, Rauth AM, Li Y, Wu XY (2007) Chemotherapy with anticancer drugs encapsulated in solid lipid nanoparticles. Adv Drug Deliv Rev 59: 491-504. [Crossref]
- Allen WL, Coyle VM, Johnston PG (2006) Predicting the outcome of chemotherapy for colorectal cancer. Curr Opin Pharmacol 6: 332-336. [Crossref]
- Zhang N, Yin Y, Xu SJ, Chen WS (2008) 5-Fluorouracil: mechanisms of resistance and reversal strategies. Molecules 13: 1551-1569. [Crossref]
- Weiner AL. Liposomes as carriers for polypeptides. Advanced Drug Delivery Reviews,.3: 307-341.
- Allen TM, Cullis PR (2013) Liposomal drug delivery systems: from concept to clinical applications. Adv Drug Deliv Rev 65: 36-48. [Crossref]
- Karanth H, Murthy RS (2007) pH-sensitive liposomes--principle and application in cancer therapy. J Pharm Pharmacol 59: 469-483. [Crossref]
- Li L, ten Hagen TL, Hossann M, Suss R, van Rhoon GC, Eggermont AM, et al. Mild hyperthermia triggered doxorubicin release from optimized stealth thermosensitive liposomes improves intratumoral drug delivery and efficacy. J Control Release. 2013;168(2): 142-150.
- Sen K, Mandal M (2013) Second generation liposomal cancer therapeutics: transition from laboratory to clinic. Int J Pharm 448: 28-43. [Crossref]
- Glavas-Dodov M, Fredro-Kumbaradzi E, Goracinova K, Simonoska M, Calis S, et al. (2005) The effects of lyophilization on the stability of liposomes containing 5-FU. Int J Pharm 291: 79-86. [Crossref]
- Luo Q, Wang P, Miao Y, He H, Tang X. A novel 5-fluorouracil prodrug using hydroxyethyl starch as a macromolecular carrier for sustained release. Carbohydrate Polymers. 2012;87(4): 2642-2647.
- Dicheva BM, ten Hagen TL, Schipper D, Seynhaeve AL, van Rhoon GC, Eggermont AM, et al. (2014) Targeted and heat-triggered doxorubicin delivery to tumors by dual targeted cationic thermosensitive liposomes. J Control Release 195: 37-48.
- Anupama Munshi, Marvette Hobbs, Meyn RE (2005) Clonogenic Cell Survival Assay. Chemosensitivity: InVitro Assays: 1.
- Zhang H, Tang J, Li C, Kong J, Wang J, et al. (2015) MiR-22 regulates 5-FU sensitivity by inhibiting autophagy and promoting apoptosis in colorectal cancer cells. Cancer Lett 356: 781-790. [Crossref]
- Ashwanikumar N, Kumar NA, Nair SA, Kumar GS (2014) Dual drug delivery of 5-fluorouracil (5-FU) and methotrexate (MTX) through random copolymeric nanomicelles of PLGA and polyethylenimine demonstrating enhanced cell uptake and cytotoxicity. Colloids Surf B Biointerfaces 122: 520-528.
- Kim IY, Kang YS, Lee DS, Park HJ, Choi EK, et al. (2009) Antitumor activity of EGFR targeted pH-sensitive immunoliposomes encapsulating gemcitabine in A549 xenograft nude mice. J Control Release 140: 55-60. [Crossref]
- Zhang J (2004) Pharmaco attributes of dioleoylphosphatidylethanolamine/ cholesterylhemisuccinate liposomes containing different types of cleavable lipopolymers. Pharmacological Research 49: 185-198.
- Eloy JO, Claro de Souza M, Petrilli R, Barcellos JP, Lee RJ, et al. (2014) Liposomes as carriers of hydrophilic small molecule drugs: strategies to enhance encapsulation and delivery. Colloids Surf B Biointerfaces 123: 345-363. [Crossref]
- Bellezza F, Alberani A, Nocchetti M, Marsili V, Cipiciani A (2014) Intercalation of 5-fluorouracil into ZnAl hydrotalcite-like nanoparticles: Preparation, characterization and drug release. Applied Clay Science 101: 320-326. [Crossref]
- Kal HB, Barendsen GW (1973) Correspondence: radiosensitivity of surviving cells in tumours pretreated with continuous irradiation. Br J Radiol 46: 1083. [Crossref]




 2021 Copyright OAT. All rights reserv
2021 Copyright OAT. All rights reserv




