Abstract
Aims: We identified the high-risk HPV genotypes associated with high-grade dysplastic cervical lesions in Burkina Faso. The available vaccines to Burkina Faso only protect against two high risk HPV genotypes: HPV 16 and 18. Are the genotypes identified in the high-grade precancerous lesions in this survey, covered by the available vaccines?
Methods: The detection and genotyping of high-risk HPV have been conducted based on 118 formalin-fixed and paraffin-embedded archived tissues using a kit named “HPV Genotypes 14 Real-TM Quant” (Sacace biotechnologies®, Italy) allowing the detection of fourteen high-risk HPV genotypes: HPV 16, 31, 18, 39, 45, 59, 33, 35, 56, 68, 51, 52, 58 and 66).
Results: The prevalence of high-risk HPV infections was 48.8% based on the appropriate PCR results (21/43). The most common HPV genotypes were HPV 39 (21.7%), HPV 35 (13.0%) and HPV 45 (13.0%). Two cases of multiple infections between HPV 39-45 and HPV 39-59 have been observed. HPV 16 was not detected in this study.
Conclusions: we noted a high prevalence rate for HPV 39, HPV 35 and HPV 45, which are not covered by the commercial vaccines. We also found that the prevalence of HPV 18 is one of the lowest prevalence rates of this study and HPV 16 is simply missing.
Key words
HPV, cancer, CIN, cervix, Burkina Faso
Introduction
Human Papillomavirus (HPV) are involved in cervical cancer, cancers of the anogenital and oropharyngeal areas, benign lesions such as condylomata acuminata and warts [1,2]. It is the most common sexually transmitted infection (STI) in the world with 660 million people infected according to the World Health Organization [3]. In Sub-Saharan Africa, cervical cancer is the first cancer in women and the most common cause of cancer deaths [4]. Developing countries bear the heaviest burden with 80% mortality [3,5]. Cervical cancer occurrence is induced by High-risk Human Papillomavirus (HR-HPV). Precancerous lesions precede cancer development [2]. Prevention involves immunization and very early diagnosis of precancerous lesions whose treatment normally prevent any cancer development [1]. The two available vaccines are effective against the genotypes of HPV 6, 11, 16 and 18. Genotypes of HPV 16 and 18 are most commonly found in cases of cervical cancer in the world and particularly in Europe [6]. But this generality is not a proven certainty in Africa and particularly in Burkina Faso, which has not yet a large-scale mapping of circulating genotypes. In addition to our previous research [7-11]. This study aims to identify the HR-HPV genotypes associated with high-grade dysplastic cervical lesions in Burkina Faso.
Material and methods
This was a cross-sectional study with retrospective data collection in which all cases diagnosed between February 2009 and May 2015 as high-grade intraepithelial lesions have been selected. There were a total of 118 tissues with cases of high-grade precancerous lesions. These tissues were archived in paraffin in the Department of Anatomy and Cytopathology of the Yalgado Ouedraogo University Hospital (CHU-YO). Cuts have been made on paraffin blocks using a microtome. Samples of five thin sheets with a thickness being ≤ 20 µm were made on each cassette and stored in Eppendorf sterile and encoded tubes and transported to the Pietro Annigoni Biomolecular Research Center/Laboratory of Molecular Biology and Molecular Genetics (CERBA/LABIOGENE) for molecular testing.
DNA extraction
The HPV DNA was extracted using the commercial kit known as FFPE DNA Purification Product # 47400 (NORGEN BIOTEK CORPORATION, Canada). Before conducting the actual extraction, the samples underwent xylene-dewaxing.
The extraction was performed following the protocol provided by the manufacturer using micro-columns equipped with collecting tubes. The DNA thus extracted was stored at -20°C until amplification.
Detection of HR-HPV genotypes through real-time PCR
Amplification was done using the kit named HPV Genotypes 14 Real-TM Quant (Sacace Biotechnologies®, Italy) which is a real-time multiplex PCR test for the detection of 14 HR-HPV genotypes (HPV-16 ; 18 ; 31 ; 33 ; 35 ; 39 ; 45 ; 51 ; 52 ; 56 ; 58 ; 59 ; 66 and 68). Each sample underwent multiplexed amplification in 4 tubes and each tube contained primers of the E6 and E7 target regions of three or four types of HR-HPV and human beta-globin gene as Internal Control (IC). For each sample, we prepared the following for the 4 tubes respectively: PCR-mix-1 16, 18, 31, IC; PCR-mix-1 39, 45, 59, IC; PCR-mix-1 33, 35, 56, 68; PCR-mix-1 51, 52, 58, 66. The pre-PCR steps included the following: preparing the Mix solution (PCR-buffer-FRT + Hot Start DNA Polymerase) and the Reaction Mix solution (Mix solution + each PCR-mix-1). For each sample, we introduced 15 μL of the Reaction Mix solution in the 4 tubes and we added 10 μL of the DNA extract. It was the same for the negative control and standards. The total reaction volume was 25 μL. The amplification program was: 1 cycle of 95°C for 15 min; 5 cycles of 95°C for 5 seconds followed by 60 °C for 20 seconds and 72°C for 15 seconds; and finally 40 cycles of 95°C for 5 seconds followed by 60°C for 30 seconds and 72°C for 15 seconds. The PCR is valid if the standards show signal for all the FAM, JOE, ROX, and Cy5 fluorochromes and if the negative control has no signal. The results were interpreted using the Microsoft Excel program named “HPV Genotypes 14 Real-TM.xls” (Sacace Biotechnologies®, Italy) provided by the manufacturer.
Statistical analysis and ethical considerations
The statistical analysis was conducted using SPSS Statistics 17.0. The Chi-square test was used for comparisons using the Epi Info 6.04d software. The difference was significant for p<0.05.
This study was approved by the Ethics Committee for Health Research (CERS) N°2014-8-099 of August 6, 2014. We complied with the confidentiality and anonymity condition with respect to information obtained from the various registers and the patient charts are kept strictly confidential.
Results
This study involved 118 tissues diagnosed as high-grade precancerous lesions according to the minutes of the histological result. The age of patients ranged from 22 to 74 years with a mean of 41.5 years and a standard deviation of 9.8 years.
Results of Histology and the real-time PCR
Following the histological examination, the various samples have been classified according to the Bethesda classification as grade 2 or 3 Cervical Intraepithelial Neoplasia (CIN) or CIN 2 or CIN 3. Of the samples, 46.6% presented cervicitis (acute, subacute or chronic). Samples whose β-globin internal control gene could not be amplified were classified negative β-globin. In our study, 63.6% or 75/118 samples were negative β-globin; they were therefore declared inappropriate. The histological results and the PCR results are shown in Table 1.
Table 1.Histological results and PCR.
|
|
Number
|
%
|
95% CI
|
p value
|
|
Histological Diagnostic
|
|
|
|
|
| |
CIN 2
|
32
|
27,1
|
19,3 to 36,1
|
p < 0,001
|
| |
CIN 3
|
86
|
72,9
|
63,9 to 80,7
|
|
|
Cervicites
|
55
|
46,6
|
37,4 to 56,0
|
|
|
|
|
|
|
|
|
Presence ofβ-globine
|
|
|
|
|
| |
β-globine +
|
43
|
36,4
|
27,8 to 45,8
|
p < 0,001
|
| |
β-globine -
|
75
|
63,6
|
54,2 to 72,2
|
|
|
|
|
|
|
|
Status HPV
|
|
|
|
|
| |
HPV +
|
21
|
17,8
|
11,4 to 25,9
|
p < 0,001
|
| |
HPV -
|
22
|
18,6
|
12,1 to 26,9
|
| |
Inappropriate
|
75
|
63,6
|
54,2 to 72,2
|
|
|
|
|
|
|
Total
|
118
|
100
|
Distribution of HPV genotypes
There were 21 women who were positive to at least one HR-HPV genotype out of the 118 resulting in a proportion of 17.8%. However, taking into account the results of the PCR, out of the 43 adequate results, the prevalence of HR-HPV infections is 48.8%. We will only consider the adequate results.
Frequency of the various HR-HPV genotypes found
The kit used helped to characterize 14 HR-HPV genotypes. These were HPV 16, 18, 31, 33, 35, 39, 45, 51, 52, 56, 58, 59, 66 and 68. By combining the HR-HPV genotypes, considering the fact that some women were infected with more than one genotype, the total number of genotypes found was 23 and 11 HR-HPV types have been found out of the 14 sought. Figure 1 shows the frequency of the various HPV genotypes found out of the 23 HPV genotypes.
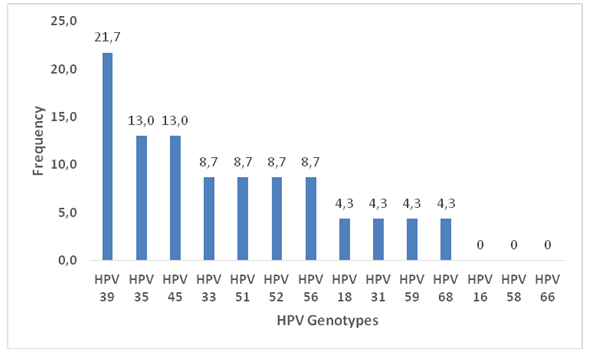
Figure 1. Frequency of HR-HPV genotypes.
The prevalence of HPV 39 infection was the highest with a frequency of 21.7%. HPV16 genotype was not detected in the study population. The prevalence of HPV 35 and HPV 45 genotypes was 13% each. Genotypes of HPV 33, 51, 52 and 56 also have the same prevalence rate of 8.7%. HPV 18, HPV 59 and HPV 68 genotypes accounted for 4.3% respectively.
Relationship between the histological diagnosis and the HR-HPV genotype
The HR-HPV genotypes found in CIN 2 represent 37.5% of the 32 cases of CIN 2 diagnosed; while HR-HPV found in CIN 3 represent only 12.8% of the 86 cases of CIN 3 diagnosed. Table 2 gives the cases of CIN 2 and CIN 3 and the corresponding HR-HPV carrier.
Table 2. HR-HPV distribution in histological lesions CIN 2 and CIN 3.
| |
Histologicaltypes
|
| |
CIN 2
|
|
CIN 3
|
|
| |
n= 32
|
%
|
95% CI
|
n = 86
|
%
|
95% CI
|
|
Genotypes HPV
|
|
|
|
|
|
|
|
HPV 16
|
0
|
0
|
0,0 to 0,0
|
0
|
0
|
0,0 to 0,0
|
|
HPV 18
|
0
|
0
|
0,0 to 0,0
|
1
|
1,2
|
0,0 to 6,3
|
|
HPV 31
|
0
|
0
|
0,0 to 0,0
|
1
|
1,2
|
0,0 to 6,3
|
|
HPV 33
|
1
|
3,1
|
0,1 to 16,2
|
1
|
1,2
|
0,0 to 6,3
|
|
HPV 35
|
1
|
3,1
|
0,1 to 16,2
|
2
|
2,3
|
0,3 to 8,1
|
|
HPV 39
|
3
|
9,4
|
2,0 to 25,0
|
2
|
2,3
|
0,3 to 8,1
|
|
HPV 45
|
3
|
9,4
|
2,0 to 25,0
|
0
|
0
|
0,0 to 0,0
|
|
HPV 51
|
1
|
3,1
|
0,1 to 16,2
|
1
|
1,2
|
0,0 to 6,3
|
|
HPV 52
|
0
|
0
|
0,0 to 0,0
|
2
|
2,3
|
0,3 to 8,1
|
|
HPV 56
|
1
|
3,1
|
0,1 to 16,2
|
1
|
1,2
|
0,0 to 6,3
|
|
HPV 58
|
0
|
0
|
0,0 to 0,0
|
0
|
0
|
0,0 to 0,0
|
|
HPV 59
|
1
|
3,1
|
0,1 to 16,2
|
0
|
0
|
0,0 to 0,0
|
|
HPV 66
|
0
|
0
|
0,0 to 0,0
|
0
|
0
|
0,0 to 0,0
|
|
HPV 68
|
1
|
3,1
|
0,1 to 16,2
|
0
|
0
|
0,0 to 0,0
|
|
Total
|
12
|
37,5
|
|
11
|
12,8
|
|
Distribution of HPV genotypes in multiple infection cases
The two cases of multiple infections are carried by women aged 31 and 40 years. They represented 9.5% against 90.5% for simple infections. These two associations were HPV 39; 45 and HPV 39; 59. The HPV 39 genotype which is the most represented genotype was present in 100% of multiple infection cases.
Distribution of the HR-HPV carrier according to age groups
HPV 39 genotype was the only one that is found in almost all age groups. HPV 18 genotype was found in the age group of 30-39 years.
HR-HPV infection is widespread in all age groups. However, the age groups of 30-39 years and 40-49 years were the most representative. The age group of 20-29 years is the only age group that recorded any genotype.
Table 3 shows the relationship between the histological result, the age group and the HPV genotype carrier.
Table3.Relationship between histological result, age group and HPV.
|
Histology and genotypes
|
Age group
|
Total
|
|
20 - 29
|
30 - 39
|
40 - 49
|
50 - 59
|
≥ 60
|
Unknown
|
|
CIN 2
|
HPV 33
|
0
|
0
|
1
|
0
|
0
|
0
|
1
|
|
HPV 35
|
0
|
0
|
1
|
0
|
0
|
0
|
1
|
|
HPV 39
|
0
|
1
|
1
|
0
|
1
|
0
|
3
|
|
HPV 45
|
0
|
3
|
0
|
0
|
0
|
0
|
3
|
|
HPV 51
|
0
|
1
|
0
|
0
|
0
|
0
|
1
|
|
HPV 56
|
0
|
0
|
1
|
0
|
0
|
0
|
1
|
|
HPV 59
|
0
|
0
|
1
|
0
|
0
|
0
|
1
|
|
HPV 68
|
0
|
0
|
1
|
0
|
0
|
0
|
1
|
|
Total
|
0
|
5
|
6
|
0
|
1
|
0
|
12
|
|
CIN 3
|
|
|
|
|
|
|
|
|
|
HPV 18
|
0
|
1
|
0
|
0
|
0
|
0
|
1
|
|
HPV 31
|
0
|
0
|
0
|
0
|
0
|
1
|
1
|
|
HPV 33
|
0
|
0
|
0
|
0
|
1
|
0
|
1
|
|
HPV 35
|
0
|
0
|
2
|
0
|
0
|
0
|
2
|
|
HPV 39
|
0
|
1
|
0
|
1
|
0
|
0
|
2
|
|
HPV 51
|
0
|
1
|
0
|
0
|
0
|
0
|
1
|
|
HPV 52
|
0
|
1
|
0
|
1
|
0
|
0
|
2
|
|
HPV 56
|
0
|
0
|
1
|
0
|
0
|
0
|
1
|
|
Total
|
0
|
4
|
3
|
2
|
1
|
1
|
11
|
One limitation of our study is the small size (118) of the samples tested of formalin-fixed tissues archived in the pathology anatomy laboratory are from different services; the fixation period was not the same and this situation probably caused some samples to be inadequate or HPV negative. In addition, the use of formalin and the Bouin solution may have deteriorated the HPV DNA, hence causing the difficulties we met.
HPV positive samples were 48.8%, based on adequate results. The positivity rate is lower than most of the results according to numerous studies [12-15]. Therefore, we wonder if the quality of the samples was not the cause. Similarly, the duration of the storage of some samples could play an important role.
But many other studies have shown results that are very different from ours [13-15] In Zambia, a team [13] found a prevalence of moderate to severe dysplasia in 31% of the cases (65 HPV positive cases against 75 cases tested and 69 cases considered appropriate cases) (p <0.001). Therefore, there is a difference in our series. However, this study also had samples with the same age period as ours and there was roughly the same number of samples. However, in this study, the sample fixation area has not been discussed. Already in the 90s, Ben-Ezra et al. [16] showed that after six hours of tissue fixation by the Bouin solution, the PCR gave no result. Later, this conclusion has been questioned by a team in Brazil [17] who showed with two different protocols that it was possible to get between 33.3% and 80% positivity to PCR despite the use of the Bouin solution. This same team demonstrated that formalin-fixed tissues gave PCR positivity rates between 87.5% and 100%; and the use of formaldehyde as a fixative gave between 17.6% and 73.7% of positivity. This team extracted the RNA instead of the DNA in our case. It should also be noted that in this study, the type of the tissue was not specified.
Our HPV positivity rate was 48.8% or 21 out of 43. In Pakistan, Siddiqa et al. [14] found 94.8% (or 73/77) of HPV positive. In this study, there were samples of cancerous tissues, low-grade lesions and high-grade lesions all confused, while our samples only concerned high-grade lesions (p <0.001) indicating a significant difference of the positivity rate which is higher than that of our study. In this study, the kit could help detect 44 HPV genotypes while ours could only help to detect 14 HPV genotypes.
Among Danish women, a study found 290 adequate results over 276 cervical samples of high-grade lesions including 100% HPV positivity against 48.8% of HPV positivity in our study (p <0.001) [15]. In this study, 82.3% of samples were CIN 3 lesions and the 17.7% were CIN 2, CIN 2 to 3, and in situ adenocarcinoma, whereas in our study, there were only five adequate CIN 3 or 11.6% and only one was HPV positive.
In our study, the most frequent genotype was HPV 39 (21.7%). HPV 18 (4.3%) was less represented with only one case in the age group of 30-39 years. The notable absentee was HPV 16 genotype yet found in many studies and recognized as one of the highest prevalence rates and being the subject of prophylactic vaccines currently available. In South Africa, McDonald et al. [18] showed high proportion of cervical lesions (CIN2 and CIN3, respectively) due to HPV 16 (22.58% and 40.28%), HPV 35 (31.18% and 19.44%) and HPV 33 (12.90 % and 12.50%). However, the team found that the genotypes of HPV 56, 59, 51 and 68 were uncommon in lesions of CIN 2 to 3. It should be noted that we included only women with a negative HIV status in this study.
Among Cameroonian women with invasive cervical cancer, the three most common high-risk types were HPV 16 (88%), HPV 45 (32%) and HPV 18 (14.8%) [19]. In New Zealand, Simonella et al. [20] showed that HPV 16 was the most represented followed by HPV 52, 31, 33, 18 HPV 39 which is nevertheless predominant in our study only represented 9.2% in CIN 2 and 5.4% in CIN 3. In general and especially in studies in developed countries, we find that the most common genotypes are HPV 16, HPV 31, HPV 52, HPV 33 and HPV 18. In Burkina Faso, the different studies produced results that are different from those found in these countries. A study conducted among HIV-positive women showed that the dominant HPV genotypes were those of the 50’S family, followed by HPV 18, the 30’S family, HPV 16 and HPV 6 [7]. The team of Djigma et al. [8] with a population of 250 HIV-positive women, frequently found genotypes of the 50’S family closely followed by HPV 18, the 30’S family, HPV 16, HPV 6, HPV 45 and HPV 11. In a female population attending gynecological consultation, Ouedraogo et al. [9] found that the most common genotype was from the 50’S family in 31% of cases. It was followed by HPV 18, HPV 16, the 30’S family, HPV 45, HPV 6 [9]. The predominant genotypes in these studies were those of the 50’S and 30’S families as well as genotypes 18 and 16. However, it should be noted that the HPV detection kit used by these authors did not allow them to accurately specify the genotypes. By characterizing these samples, Zohoncon et al. [10] found HPV 52 followed by HPV 18, 31 and 35, while genotypes of HPV 59 and 33 were the less frequent. In a study conducted among teen girls in Ouagadougou, Ouedraogo et al. [9] found like Zohoncon et al. [10], that HPV 52 was the most common genotype with 22.8%, followed by genotypes of HPV 59, HPV 39, HPV 35 and HPV 51 [11]. In these studies, the serological status of most patients as well as the socio-demographic characteristics were all known; which is different in our series.
We encountered two cases of multiple infections (double HPV) in 9.5% of the cases. Garcia-Espinosa et al. [18] found 18.4% of multiple infections [21]. In South Africa, 24% of cases of multiple infections were observed. In Burkina Faso, 78.03% of cases of multiple infections were observed, but 90.1% of the patients were HIV-infected [10]. Several cases of multiple infections were found by Ouedraogo et al. [11]; in this case, the number of HPV genotypes present in the 35 cases of multiple infections was 88 out of a total of 136 genotypes (64.7%) and related to 42.2% of infected women. HPV 52 accounted for 23.8% of genotypes present in these multiple infections (21/88); there were associations of 2 to 5 HPV types. However, in our study, we found that the HPV 39 genotype was present in both cases of dual infections representing 100%. A study in South Africa found that HIV-positive women frequently showed associations of over two genotypes [22]. This could explain the fact that the studies which included HIV-infected women showed more multiple infections.
We did not observe any correlation between the different associations of HPV genotypes, as the phenomenon is not well understood. We found no significant link between the histological lesions and the HPV carrier. In our study, the prevalence rate of precancerous lesions was significantly lower than that of most studies [13,21,15]. This could be explained by the size of our sample but especially by the quality of our samples; or other factors other than HPV such as bacteria, parasites, fungi, other virus, or simply other HPV genotypes that our detection kit could not characterize.
The size of our sample is not large enough to properly assess the relationship between the age and the HPV carrier. The higher age in our study could be the reason why we did not find some genotypes due to clearance; this could also explain the low HPV prevalence.
Conclusion
The results of our study show that the distribution of the HPV types can be determined by using pathological anatomy tissues such as paraffin blocks which allow the assessment of changes over time. But precautions shall be taken to avoid false negatives and inadequate samples. However, we should optimize the methods of histology sample conservation in order to keep the DNA of these cells as intact as possible. Cervical cancer can be prevented by early diagnosis of precancerous lesions and through molecular characterization of the various genotypes circulating in each geographical area in order to better direct research on new multivalent vaccines. To do this, the establishment of an African mapping of high-risk HPV is required.
Acknowledgments
The authors are thankful to the Pietro ANNIGONI Association and the WAEMU Commission through the PACER-II program for their financial support.
Conflicts of Interest notification
All authors declare there is no conflict of interest in this work.
References
- Monsonégo J (2006) Cervical cancer prevention: the impact of HPV vaccination. GynecolObstetFertil34: 189-201. [Crossref]
- Monsonego J (2010) Anal cancer and human papillomaviruses: a perspective based on the cervical cancer model. GynecolObstetFertil 38: 250-254. [Crossref]
- World Health Organization (2007) The fight against cervical cancer. Essential Practices Guide. Integrated support for reproductive and sexual health and chronic disease. Genève 287.
- Anorlu RI (2008) What is the significance of the HPV epidemic? Can J Urol 15: 3860-3865. [Crossref]
- Ly A (2009) Le cancer du col utérin: nouveaux vaccins, nouvelles perspectives? J Afr Cancer 1: 65-67.
- Heard I, Tondeur L, Arowas L, Falguières M, Demazoin MC, et al. (2013) Human papillomavirus types distribution in organised cervical cancer screening in France. PLoS One 8: e79372. [Crossref]
- Sagna T, Djigma F, Zeba M, Bisseye C, Karou SD, et al. (2010) Human papillomaviruses prevalence and genital co-infections in HIV-seropositive women in Ouagadougou (Burkina Faso). Pak J BiolSci 13: 951-955. [Crossref]
- Djigma FW, Ouédraogo C, Karou DS, Sagna T, Bisseye C, et al. (2011) Prevalence and genotype characterization of human papillomaviruses among HIV-seropositive in Ouagadougou, Burkina Faso. Acta Trop 117: 202-206. [Crossref]
- Ouedraogo CM, Djigma FW, Bisseye C, Sagna T, Zeba M, et al. (2011) Epidemiology, characterization of genotypes of human papillomavirus in a population of women in Ouagadougou. J GynecolObstetBiolReprod (Paris) 40: 633-638. [Crossref]
- Zohoncon TM, Bisseye C, Djigma FW, Yonli AT, Compaore TR, et al. (2013) Prevalence of HPV High-Risk Genotypes in Three Cohorts of Women in Ouagadougou (Burkina Faso). Mediterr J Hematol Infect Dis 5: e2013059. [Crossref]
- Ouédraogo CM, Rahimy RM, Zohoncon TM, Djigma FW, Yonli AT, et al. (2015) Epidemiology and characterization of high-risk genotypes of human Papillomavirus in a population of sexually active adolescents in Ouagadougou. J GynecolObstetBiolReprod (Paris) 44: 715-722. [Crossref]
- Hammer A, Mejlgaard E, Gravitt P, Høgdall E, et al. (2015) HPV genotype distribution in older Danish women undergoing surgery due to cervical cancer. ActaObstetGynecolScand 94: 1262-1268. [Crossref]
- Bateman AC, Katundu K, Polepole P, Shibemba A, Mwanahamuntu M, et al. (2015) Identification of human papillomaviruses from formalin-fixed, paraffin-embedded pre-cancer and invasive cervical cancer specimens in Zambia: a cross-sectional study. Virol J 12: 2. [Crossref]
- Siddiqa A, Zainab M, Qadri I, Bhatti MF, Parish JL (2014) Prevalence and genotyping of high risk human papillomavirus in cervical cancer samples from Punjab, Pakistan. Viruses 6: 2762-2777. [Crossref]
- Kirschner B, Schledermann D, Holl K, Rosenlund M, Raillard A, et al. (2013) HPV-genotypes in high-grade intraepithelial cervical lesions in Danish women. ActaObstetGynecolScand 92: 1032-1040. [Crossref]
- Ben-Ezra J, Johnson DA, Rossi J, Cook N, Wu A (1991) Effect of fixation on the amplification of nucleic acids from paraffin-embedded material by the polymerase chain reaction. J HistochemCytochem 39: 351-354. [Crossref]
- Gouveia GR, Ferreira SC, Ferreira JE, Siqueira SA, Pereira J (2014) Comparison of two methods of RNA extraction from formalin-fixed paraffin-embedded tissue specimens. Biomed Res Int 2014: 151724. [Crossref]
- McDonald AC, Tergas AI, Kuhn L, Denny L, Wright TC Jr (2012) Distribution of high-risk human papillomavirus genotypes among HIV-negative women with and without cervical intraepithelial neoplasia in South Africa. PLoS One 7: e44332. [Crossref]
- Pirek D, Petignat P, Vassilakos P, Gourmaud J, Pache JC, et al. (2015) Human papillomavirus genotype distribution among Cameroonian women with invasive cervical cancer: a retrospective study. Sex Transm Infect 91: 440-444. [Crossref]
- Simonella LM, Lewis H, Smith M, Neal H, Bromhead C, et al. (2013) Type-specific oncogenic human papillomavirus infection in high grade cervical disease in New Zealand. BMC Infect Dis 13: 114. [Crossref]
- García-Espinosa B, Moro-Rodríguez E, Alvarez-Fernández E (2012) Genotype distribution of human papillomavirus (HPV) in histological sections of cervical intraepithelial neoplasia and invasive cervical carcinoma in Madrid, Spain. BMC Cancer 12: 533. [Crossref]
- Adler D, de Bruyn G, Laher F, Gray G, Allan BR, et al. (2008) Human papillomavirus genotype distribution among human immunodeficiency virus (HIV)-infected and non-HIV-infected women in Soweto, South Africa. J ClinMicrobiol46: 4109-4110. [Crossref]

