Prostate cancer appears in two versions: familial and sporadic. In the development of sporadic prostate cancer, sequence variations in some genes, especially in genes involved in steroid hormone metabolism has been considered important. CYP19 gene is one of the most interesting. The product of this gene converts the C19 steroid to estrogen. Some studies have shown long allele of TTTA in intron4, as a tetra nucleotide repeat is associated with increased risk of prostate cancer.In this case/ control research we studied the association of TTTA repeat variations and prostate cancer risk among 59 patients with sporadic prostate cancer and 95 healthy men which were referred to Shohada-e -Tajrish Hospital, under SBMU license. This study did not revealed significant association between various length of tetra nucleotide repeats (including long and short repeats) and prostate cancer risk (with P-value=0.4). Results of our study do not support about considered role of TTTA repeat variations and prostate cancer in Iranian population.
iranian population, CYP19, TTTA repeat variations, prostate cancer
In view of the recent studies, prostate cancer incidence has been reported 9.6 men (3.2-16) per 100,000 in Iranian population [1] and is considered as a major cause of mortality among men [2]. This information indicates the importance of study in this area for health improvement in Iran.
Sequence variation in members of CYP450 gene super family is considered as one of the most important genetic susceptibility factor for sporadic prostate cancer. Among, association of CYP19 polymorphism (rs60271534), TTTA repeat length variations, has been studied in many population and many of these have shown a significant association between long allele (over than seven repeats) and increased risk of prostate cancer [3] and also with poor clinical prognosis [4]. Furthermore, one research has been appeared similar results regarding breast cancer [5].
CYP19 gene is located 15q21.1 and has 123,000 base pairs and 10 exons. This gene is transcribed from telomere to centromere [6,7]. Product of gene plays a role in C19 steroid conversion to estrogen. It seems that the activity of produced enzyme has been increased in some prostate cancer tissues [8]. The variation in question occupies intron 4. One research has been shown that the high-repeat genotype (>nine repeats) is associated with high level estrogen [9]. But, by the now, the exact role of this polymorphism has not been clearly known.
Patients
In this research, we studied 154 unrelated Iranian referred men to the Shohada-e- Tajrish hospital under SBMU license, Tehran, Iran, including 59 patients with prostate cancer (case group) and 95 healthy men (control group). All subjects were of Iranian ethnogeographic origin. Two groups were matched for age. Selection of cases was done based on positive pathology result. Control samples were selected based on normal result of DRE (Digital Rectal Examination) and PSA levels less than 4 ngr/milt [10]. Age range of people in two groups was 50 to 75 [11]. People attending in this research had not family history of prostate cancer. Also, we acquired some information about pathologic features of tumor tissues via pathology report sheets.We gathered information about smoking and opiate addiction with questionnaire, too.In this study all of people participated with informed consent.
Genotyping
Peripheral blood samples were gathered in K3EDTA tube. Extraction of DNA was done with "Diatom TM DNA Prepkit" (Diatom, Russia). Then PCR was carried out with 25 μl as a total volume and components including: 1.5 mmol 10X PCR buffer with 160 mM (NH<4)2SO4 -670mM Tris-HCl - 0.1% tween-20 contain (KBC, Iran), 1.5mmol Mgcl2(KBC, Iran), 200 μmol dNTPs (Bioscience, England), 8pmol from each primers (Bioneer, South Korea), 2.5IU Taq pol (KBC, Iran) and 200 ngr of genomic DNA. The sequence of primers which was described by Tsuchiya et al. has been indicated in Table 1.
Table 1. Sequences of Forward and Reverse Primer
Forward primer |
5ꞌ-TTATGAAAGGTAAGCAGGTACTTAG-3ꞌ |
Reverse primer |
5ꞌ-GTCGTGAGCCAAGGTCACT-3ꞌ |
PCR was done, respectively: with initial denaturation 95°C in 6 minute and 35 cycles including: denaturation step 94°C 40s, annealing step 54.5°C 45s, replication step 72°C 45s and final replication 72°C 6 min. PCR products were studied with 10% polyacrylamide gel electrophoresis (MERCK, Germany), (165 min, 200V). Then the gels were stained with silver nitrate (Ghatran chimi, Iran) and observed with white light. In this stage, samples were checked about number of bands and covered distance along the gel.Subsequently, some of homozygote samples were selected and studied with sequencing. Then, samples with seven repeats were used as a ladder to determine the approximate number of repeats of other samples on 10% PAGE.
Statistical analysis
Unprocessed data were analyzed with T-test and chi-square test program of SPSS. Association between each of genotypes ((TTTA)n) , probable risk factors and prostate cancer riskwas assessed by calculating P-value with 95% confidence intervals. P-value<0.05 was considered significant.
Mean age of participants in this study was 68.46 and 68.16 for cases and controls, respectively with no significant difference between them (P-value=0.830). Also, we studied the PSA level in patients and healthy men, too. This study indicated significant difference between two groups. Mean of PSA levels in cases and controls was 15.43 ngr/milt and 1.92 ngr/milt (P-value<0.0001, CI=95%).
As well, the Gleason score at three groups including low ≤4 and moderate 4-6 and high score ≥7 was compared in patients. There was no significant difference among genotype groups (P-value>0.05).
Smoking and opiate addiction, as possible risk factors was assessed based on collected information with questionnaire. Statistical analysis revealed a significant association between smoking andincreased risk of prostate cancer, OR=4.2 (95% CI=2.0914-8.4343, P=0.0001) (Tables 2-4). As well, there was no considerable association about "smoking & opiate addiction" and prostate cancer risk, OR=3 (95% CI=0.1803_ 49.9167, P=0.4438).
An example of procedures (PAGE and sequencing) has been shown in Figure 1 and 2. Statistical analysis was done for determining association between long allele (>seven TTTA repeats) and short allele (≤seven TTTA repeats), as genotypes including S, SS, L, LL, LS, and prostate cancer risk. Relation of different genotypes and prostate cancer risk has been shown in Table 2. Results of statistical analysis did not detect an association between any genotypes and prostate cancer with P-value=0.4 in cases and P-value=0.4 in controls. In addition, there was no significant difference about long allele and short allele between case and control groups OR=1.0551 (95% CI= 0.5763- 1.9319, P=0.8619).
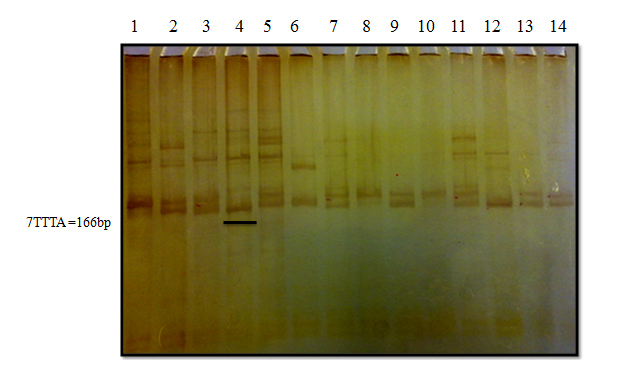
Figure 1. Example of 10% PAGE. Sample with seven repeats (lane 4) was used as a ladder to determine length of other samples.
≤7: short allele, >7: long allele
Genotypes were included:
S (homozygote with mass short alleles), SS (heterozygote with different short alleles)
L (homozygote with similar long alleles), LL (heterozygote with different long alleles)
SL (heterozygote with short and long alleles)
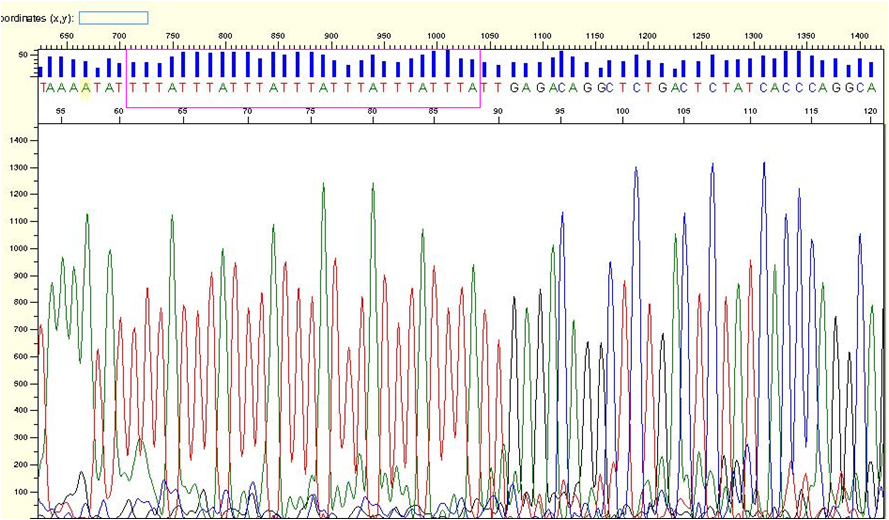
Figure 2. Example of sequencing. This sample shows seven TTTA repeat. Sample with seven repeats was used as a ladder to determine the number of TTTA repeats about other samples. (With Saq Scanner software)
Table 2. Relation of different genotypes of CYP19 and prostate cancer risk
|
|
Total |
Asymp. Sig. (2 sided) P value |
S |
SS |
L |
LL |
LS |
Groups |
Case |
Count
% within Groups |
8
13.6% |
8
13.6% |
10
16.9% |
23
39.0% |
10
16.9% |
59
100.0% |
.400 |
Control |
Count
% within Groups |
20
21.1% |
7
7.4% |
11
11.6% |
35
36.8% |
22
23.2% |
95
100.0% |
.400 |
Total |
Count
% within Groups |
28
18.2% |
15
9.7% |
21
13.6% |
58
37.7% |
32
20.8% |
154
100.0% |
.949 |
*P-value<0.05 is significant
S: short allele (homozygote); SS: short allele (heterozygote); L: long allele (homozygote); LL: long allele (heterozygote); LS: long allele/short allele ≤7 TTTA: short allele; >7TTTA: long allele
Table 3. Statistical result of relation between Gleason Score and types of genotype
|
CYP19 |
Total |
Asymp. Sig. (2 sided) P value |
|
|
|
S |
SS |
L |
LL |
LS |
GLEASON Score |
1 |
Low
% within GLEASON |
0
0% |
0
0% |
0
0% |
2
100.0% |
0
0% |
2
100.0% |
.271 |
|
2 |
Moderate
% within GLEASON |
3
12.5% |
5
20.8% |
5
20.8% |
10
41.7% |
1
4.2% |
24
100.0% |
.191 |
|
3 |
High
% within GLEASON |
5
15.6% |
3
9.4% |
4
12.5% |
11
34.4% |
9
28.1% |
32
100.0% |
.456 |
Total |
|
Count
% within GLEASON |
8
13.8% |
8
13.8% |
9
15.5% |
23
39.7% |
10
17.2% |
58
100.0% |
|
*P-va1ue<0.05 is significant
Table 4. Relation between addiction and prostate cancer risk
|
|
|
Groups |
Total |
P-value |
|
|
|
Case |
Control |
Addiction |
smoking |
Count
% within Groups |
35
59.32% |
25
26.31% |
60
38.96% |
0.0001 |
|
Smoking and Opiate |
Count
% within Groups |
1
1.69% |
1
1.05% |
2
1.29% |
0.4438 |
|
No addiction |
Count
% within Groups |
23
38.98% |
69
72.63% |
92
|
0.0001 |
Total |
|
Count
% within Groups |
58
100% |
95
100% |
154
100% |
|
*P-va1ue< 0.05 is significant
Prostate cancer may appear in familial or sporadic version. Familial type, with low prevalence, is attributed to mutations in HPC1, HPC2 locus, MSR1 gene and other genes with high penetrance [12,13]. But, sporadic prostate cancer has high prevalence. In this type of prostate cancer, mutations or sequence variations in low penetrance genes, such as CYP19, other members of CYP450 gene superfamily and so are considered. In some studies, similar to our study, analysis of TTTA repeat variations in CYP19 and prostate cancer risk, have been done considering seven repeats of TTTA as the cut off [4]. In one study, association of this polymorphism and breast cancer risk has been carried out for assessing the exact number of repeats, exactly (less than seven, 7, 8, 9, 10, 11, 12, 13 repeats) [5]. The used technique in recent study has been "gene scan". But, we had restriction about used technique and we had to do procedure which has been explained in "Materials and Methods". Although, some studies have shown significant difference about genotype groups of TTTA polymorphism between case and control groups [14]. For example, Latil et al. showed 171 and 187 allele association with prostate cancer risk (P-value=0.05 and 0.045, respectively). Also, Tsuchiya et al. indicated TTTA>7 repeat is associated with a worth clinical prognosis, in patients with Japanese origin (P-value=0.016). But, our research did not appear any distinctness among cases and controls (P-value=0.4). We did not observe differences aboutpathology characteristics between types of genotype in patients, too. Therefore, this polymorphism may not influence prostate cancer development in Iranian men, likely. Considering Iranian ethnicity of participated people, this study results are not susceptible to population stratification bias [15-19].
In conclusion, our study did not detectan association between CYP19 (TTTA)n repeat polymorphism and risk or aggressivenessof prostate cancer. But, smoking could be considered as a probable environmental risk factor for sporadic prostate cancer. Although, our research did not show significant difference about genotypes of studied polymorphism, but these results and other studies could be used for understanding the pathophysiology in sporadic prostate cancer.
We thank the "student research committee" of Shahid Beheshti University of Medical Science and also, Dr Vahhid Reza Yassaee, Miss Nilufar Safavi andMr.Zarrin baf, head nurse of Shohada-e- Tajrish hospital urology department. The data provided in this article was extracted from the Master Degree thesis of Arezu Karimpur. The thesis was conducted under supervision of Dr Farkhondeh Pouresmaili, Dr Farah Farzaneh and Dr Movafagh at the Department of medical genetics, Shahid Beheshti University of Medical Sciences, Tehran, Iran.
- 1. Alberti C (2010) Hereditary/familial versus sporadic prostate cancer: few indisputable genetic differences and many similar clinicopathological features. Eur Rev Med Pharmacol Sci 14: 31-41. [Crossref]
- 2. Baxter SW, Choong DY, Eccles DM, Campbell IG (2001) Polymorphic variation in CYP19 and the risk of breast cancer. Carcinogenesis 22: 347-349. [Crossref]
- 3. Camp NJ, Tavtigian SV (2002) Meta-analysis of associations of the Ser217Leu and Ala541Thr variants in ELAC2 (HPC2) and prostate cancer. Am J Hum Genet 71: 1475-1478. [Crossref]
- 4. Chen H, Griffin AR, Wu YQ, Tomsho LP, Zuhlke KA, et al. (2003) RNASEL mutations in hereditary prostate cancer. J Med Genet 40: e21. [Crossref]
- 5. Cussenot O, Azzouzi AR, Nicolaiew N, Fromont G, Mangin P, et al. (2007) Combination of polymorphisms from genes related to estrogen metabolism and risk of prostate cancers: the hidden face of estrogens. J Clin Oncol 25: 3596-3602. [Crossref]
- 6. Gennari L, Masi L, Merlotti D, Lucia Picariello, AlbertoFalchetti, et al. (2004) A polymorphic CYP19 TTTA repeat influences aromatase activity and estrogen levels in elderly men: effects on bone metabolism. J Clini Endocrinol Metab 89: 2803-2810. [Crossref]
- 7. Gsur A, Feik E, Madersbacher S (2004) Genetic polymorphisms and prostate cancer risk. World J Urol 21: 414-423. [Crossref]
- 8. Haiman CA, Hankinson SE, Spiegelman D, Brown M, Hunter DJ (2002) No association between a single nucleotide polymorphism in CYP19 and breast cancer risk. Cancer Epidemiol Biomarkers Prev 11: 215-216. [Crossref]
- 9. Hamilton SR, Aaltonen LA (2000) Pathology and genetics of tumours of the digestive system. IARC press Lyon.
- 10. Hiramatsu M, Maehara I, Ozaki M, Harada N, Orikasa S, et al. (1997) Aromatase in hyperplasia and carcinoma of the human prostate. Prostate 31: 118-124. [Crossref]
- 11. Hong WK, Hait WN, Kufe D, Kufe DW, Holland JF, et al. (2010) Holland Frei cancer medicine eight. PMPH-USA.
- 12. Latil AG, Azzouzi R, Cancel GS, Guillaume EC, Cochan-Priollet B, et al. (2001) Prostate carcinoma risk and allelic variants of genes involved in androgen biosynthesis and metabolism pathways. Cancer 92: 1130-1137. [Crossref]
- 13. Li L, Cicek MS, Casey G, Witte JS (2004) No association between a tetranucleotide repeat polymorphism of CYP19 and prostate cancer. Cancer Epidemiol Biomarkers Prev 13: 2280-2281. [Crossref]
- 14. Mousavi SM (2009) Toward prostate cancer early detection in Iran. Asian Pac J Cancer Prev 10: 413-418. [Crossref]
- 15. Pourmand G, Salem S, Mehrsai A, Lotfi M, Amirzargar MA, et al. (2007) The risk factors of prostate cancer: a multicentric case-control study in Iran. Asian Pac J Cancer Prev 8: 422-428. [Crossref]
- 16. Sadjadi A, Nooraie M, Ghorbani A, Alimohammadian M, Zahedi MJ, et al. (2007) The incidence of prostate cancer in Iran: results of a population-based cancer registry. Arch Iran Med 10: 481-485. [Crossref]
- 17. Sarma AV, Dunn RL, Lange LA, Ray A, Wang Y, et al. (2008) Genetic polymorphisms in CYP17, CYP3A4, CYP19A1, SRD5A2, IGF-1, and IGFBP-3 and prostate cancer risk in African-American men: the Flint Men's Health Study. Prostate 68: 296-305. [Crossref]
- 18. Severi G, Morris HA, MacInnis RJ, English DR, Tilley W, et al. (2006) Circulating steroid hormones and the risk of prostate cancer. Cancer Epidemiol Biomarkers Prev 15: 86-91. [Crossref]
- 19. Tsuchiya N, Wang L, Suzuki H, Segawa T, Fukuda H, et al. (2006) Impact of IGF-I and CYP19 gene polymorphisms on the survival of patients with metastatic prostate cancer. J Clin Oncol 24: 1982-1989. [Crossref]


