Abstract
To explore how screening for potential nutrient bio-activity could be improved by integrating biological and chemical markers of cellular responses, we assessed the effect of the known anti-oxidants, catechin and chlorogenic acid, on anti-oxidant activity in vitro. We determined an active concentration (25 µM) of each compound and then measured its effects on anti-oxidant gene expression and regulation, enzyme activity, and cellular anti-oxidant activity. We found that only catechin retained biological anti-oxidant effects in vitro, although catechin and chlorogenic acid are chemical anti-oxidants. A limited role for microRNA regulation of anti-oxidant was found, however it cannot be excluded as a possible mechanism because other microRNAs and/or epigenetic factors may be involved. Our approach found that screening for bio-activity of nutrients, in this example anti-oxidant activity, is more appropriate using an integrated approach and highlights the importance of using a range of techniques, such as those used in this study, to properly characterise the effects of dietary ingredients.
Key words
epigenetics, microRNA, oxidative stress, phytochemical
Introduction
Our understanding of the effects of nutrients on cellular function requires appropriate in vitro models in order to validate hypotheses of interest, or to justify further study in more complex biological models. To assess if using multiple biological measurements of cell function could improve purely chemical measurements of potential bio-activity and/or provide a more relevant biological interpretation, we developed a model to study anti-oxidant activity. Despite the recent EFSA (European Food Safety Authority) ruling that no evidence has been provided that having anti-oxidant activity/content and/or anti-oxidant properties is a beneficial physiological effect, the word anti-oxidant still resonates among consumers [1,2]. This rise in anti-oxidant awareness has enabled anti-oxidant nutrients to become a marketing tool. Although there is still debate as to what is the “gold standard” to test for nutrient benefits (and some may argue that randomised controlled trials are the ultimate test for functionality), in vitro studies like the one presented here provide a valuable tool for better mechanistic understanding.
Catechin and chlorogenic acid were used as reference dietary ingredients as they are found in many commonly consumed foods (e.g. coffee, tea, chocolate) and have been shown to act as potent anti-oxidants in a number of studies [3-14]. Epidemiological studies have suggested a link between the consumption of foods rich in these compounds and reduced risk of disease [15-17]. However, most of the preliminary biological studies investigating the capacity of these compounds, or their metabolites, to affect the cellular response to oxidative damage have used highly transformed cancer cell models. How catechin and chlorogenic acid affect the balance of reactive oxygen species (ROS) regulation by the CAT, SOD, and GPX pathways in less diseased or normal cells is not known, nor is the effect of these compounds on the activity of miRNAs regulating anti-oxidant gene expression [18]. ROS are involved in several physiological processes and an imbalance in their abundance may contribute to the risk of disease [19]. The balance of ROS levels in cells is controlled by the catalase (CAT), superoxide dismutase (SOD), and glutathione peroxidase (GPX) anti-oxidant pathways [20-26]. There has been considerable interest in recent years in assessing how phytochemicals from our diet could affect the ROS balance within cells to reduce inappropriate ROS activity [27-32].
However, these studies have largely focused on the chemical anti-oxidant activity of these compounds which often requires a non-physiological pH and temperature; very few of these studies take into account the bioavailability, uptake, and metabolism of the anti-oxidant compounds. Biological systems are much more complex than the simple chemical mixtures employed, and anti-oxidant compounds may operate via multiple mechanisms.An understanding of how potential anti-oxidants from food affect cellular anti-oxidant pathways in normal or early stage disease models is essential in assessing possible health benefits.
The expression and activity of the CAT, SOD1-3, and GPX1-8 genes (and resulting capacity to manage intracellular ROS levels) are influenced by several factors such as mutations in the coding sequence or defective protein folding, both of which lead to inappropriate anti-oxidant
function [20,22,24,30]. However, the role of epigenetic factors such as microRNA in these pathways, and how phytochemicals may influence these epigenetic factors, is poorly understood [33,34]. MicroRNAs (miRNAs) are post-transcriptional regulators that profoundly co-ordinate cell physiology by influencing mRNA stability leading to translational repression of protein-coding genes. The ability of miRNAs to regulate many distinct mRNA transcripts and the evidence for inappropriate miRNA regulation in disease suggests a fundamental role for miRNAs in how cell biology changes during disease [35]. Furthermore, dietary polyphenols, for example catechin, have been shown to influence the expression of miRNAs and affect markers of cell function and health [36-53].
The aim of the study was to develop a model to study how genetic, epigenetic, enzymatic, and phenotypic markers of the cellular response to oxidative damage may be altered by dietary factors. An immortalised (human with papillomavirus-18) prostate epithelial cell line (RWPE-1) and an early-stage prostate cancer cell line (WPE1-NA22) provided a cell model for approximately “normal” and abnormal cell health, respectively. The effects of catechin and chlorogenic acid on free radical scavenging capacity, viability, and oxidative defence, as well as expression of anti-oxidant genes and miRNAs, and anti-oxidant enzyme activity, were assessed in the RWPE-1 and WPE1-NA22 cells lines. The concentration of each compound for study was based on that which was more effective at scavenging free radicals than cell lysates alone.
Materials and methods
Cell culture
Authenticated RWPE-1 and WPE1-NA22 cells were purchased from the American Type Culture Collection (ATCC, Manassas, VA, USA). All cell culture reagents were obtained from Gibco (Life Technologies, Auckland, NZ) unless otherwise stated. Both cell lines were cultured in keratinocyte serum-free media supplemented with 25 mg of bovine pituitary extract, 2.5 µg of recombinant epidermal growth factor and 1% penicillin/streptomycin (Pen Strep). All cell lines were maintained at 37°C and 5% CO2 in a humidified atmosphere according to their individual ATCC guidelines and all experiments were completed within 12 sub-cultures from the original ATCC stock.
Chemicals
All chemicals were purchased from Sigma-Aldrich (Auckland, NZ) unless otherwise stated. Stock solutions of 2.29 mM (+)-Catechin hydrate (CAS Number 225937-10-0, ≥98%, C1251), and 3.12 mM chlorogenic acid (CAS Number 327-97-9, ≥95%, C3878) were prepared in DMSO. The negative control was culture medium supplemented with DMSO matched to the DMSO concentration in the treatment medium (0.02% (v/v)).
Free radical scavenging capacity assay
The capacity of catechin, chlorogenic acid, and untreated cell lysates to scavenge the free radical formed from the reduction of 2,2-Di(4-tert-octylphenyl)-1-picrylhydrazyl (DPPH, Sigma-Aldrich, Auckland, NZ) was measured based on changes in the absorbance (at 515 nm) of the free
radical [54-56].
For the first experiment, twelve replicates of catechin or chlorogenic acid at 0, 0.1, 1, 10 and 25 µM in methanol (Sigma-Aldrich, Auckland, NZ, HPLC grade >99.9%) were prepared and mixed with 60 µM of DPPH (in methanol) in 96-well micro-titre plates (Corning, Corning, NY, USA). After 30 minutes the absorbance of the DPPH radical was measured at 515 nm using a SpectraMax 250 spectrophotometer (Molecular Devices, Downington, PA, USA). For the second experiment, twelve lysates (using DMSO) of healthy untreated RWPE-1 or WPE1-NA22 cells (5x105 cells per replicate) were assayed as described above.
Cell viability assay
The effect of catechin or chlorogenic acid on the metabolic activity (a marker of cell viability) of the RWPE-1 and WPE1-NA22 cell lines after 24 hours was measured using the water-soluble tetrazolium (WST-1) cytotoxicity assay (Clontech, Mountain View, CA, USA). This assay is based on the cleavage of the tetrazolium salt to a formazan dye by mitochondrial succinate-tetrazolium reductase in viable cells and is quantified by measuring the formazan absorbance at 450 nm.
For each cell line, 25 µM catechin or chlorogenic acid were tested on twenty-four replicates of 9 x 103 cells per well in 96-well tissue-culture plates (Corning, Corning, NY, USA). The cells were seeded and cultured for 24 hours to facilitate attachment after which the medium was replaced with 90 µl of the appropriate supplemented (or control) medium and cultured for a further 24 hours. After this, 10 µl of WST-1 was added to each well and the plates incubated for 2 hours before the absorbance of the formazan produced was measured at 450 nm (reference 650 nm) using a SpectraMax 250 spectrophotometer (Molecular Devices, Downington, PA, USA).
Cellular oxidative defence assay
The effect of 25 µM catechin or chlorogenic acid on the cellular oxidative defence capacity of the RWPE-1 and WPE1-NA22 cell lines was measured. This assay is based on the capacity of samples to prevent the formation of dichlorofluorescein (DCF, emission at 538 nm) from dichlorofluoresceindiacetate (DCF-DA) by 2,2’-azobis(2-amidinopropane) dihydrochloride (ABAP)-generated peroxyl radicals in human cells [57].
For each cell line, 25 µM catechin or chlorogenic acid were tested on twelve replicates of 6 x 104 cells per well in opaque 96-well tissue-culture plates (Corning, Corning, NY, USA). The cells were seeded and cultured for 24 hours to facilitate attachment after which the medium was replaced with 50 µl of the appropriate supplemented (or control) medium and 50 µl of DCF-DA (Sigma-Aldrich, Auckland, NZ). After incubation (1 hour at 37°C and 5% CO2), the cells were washed with 100 µl of PBS and 100 µl of 600 µM ABAP (in Hank’s balanced salt solution, Sigma-Aldrich, Auckland, NZ) was added. The fluorescence emission at 538 nm (excitation at 485 nm) was measured every five minutes for one hour using a SpectraMax 250 spectrophotometer (Molecular Devices, Downington, PA, USA).
Quantification of oxidative defence gene expression
The expression of genes associated with oxidative defence: catalase (CAT), superoxide dismutase 2 (SOD2) and glutathione peroxidase 1 (GPX1), were quantified using TaqManreal-time PCR relative to the hypoxanthine phosphoribosyltransferase 1 (HPRT1) reference gene. RWPE-1 and WPE1-NA22 cells were exposed to 25 µM catechin or chlorogenic acid and gene expression measured at 24 hours.
For each cell line, 5 x 105 cells were seeded into sufficient wells of a 6-well tissue-culture plate (Corning, Corning, NY, USA) and cultured for 24 hours to facilitate attachment. The medium was replaced with the appropriate supplemented (or control) media and cultured for 24 hours. The medium was removed and 2.5 ml of TRI reagent (Life Technologies, Auckland, NZ) was added for
five minutes to facilitate cell lysis. After mixing the lysates were stored in sterile nucleic acid-free tubes at -80°C. The total RNA from each cell lysate was isolated using the RiboPure RNA isolation kit (Life Technologies, Auckland, NZ) according to the manufacturer’s instructions, with the following modifications; 1-bromo-3-chloropropane, 100 µl, (Sigma-Aldrich, Auckland, NZ) was used in place of chloroform, two wash steps were completed and the total RNA eluted in two volumes of 30 µl. The total RNA was stored at -80°C overnight and quantified from its A260:280 nm and A260:230 nm ratios using a NanoDrop 1000 spectrophotometer (ThermoFisher Scientific, Scoresby, Vic, AU).
For real-time PCR analysis, 900 ng of total RNA was reverse transcribed into cDNA using random octomers and a high capacity RNA-to-cDNA kit (Life Technologies, Auckland, NZ) according to the manufacturer’s instructions. The cDNA was stored at -20°C until required. The TaqMan assays (Life Technologies, Auckland, NZ) used were; CAT (Hs00156308_m1), SOD2 (Hs00167309_m1), GPX1 (Hs00829989_gH) and HPRT1 (333768F). All PCRs (no-template controls, untreated and treated samples) were prepared as triplicate 10 µl reactions comprising a 9.0 µl aliquot of master mix (5.0 µl of 2x GE master mix (Life Technologies, Auckland, NZ), 0.5 µl of 20x gene expression assay (Life Technologies, Auckland, NZ), 3.5 µl of nuclease-free water (Life Technologies, Auckland, NZ)), and 1 µl of cDNA (at a 1 in 18 dilution in nuclease-free water). The thermal profile used was; 50°C for 2 minutes, 95°C for 10 minutes and 40 cycles of 95°C for 15 seconds and 60°C for 60 seconds. The experiment was completed using a RotorGene 6000 qPCR instrument (Qiagen, Hilden, DE). The data were normalised to the reference gene and analysed for expression level changes using the ΔΔCt method. The experiment was completed as three biological replicates each with three technical replicates for all samples.
Quantification of microRNA gene expression
The expression of miRNAs predicted to target CAT, SOD2, or GPX1 mRNA translation were quantified using TaqManreal-time PCR in RWPE-1 and WPE1-NA22 cells treated with 25 µM catechin or chlorogenic acid for 24 hours The expression of the MIR107, MIR30E, and MIRLET7G genes, relative to the small nucleolar RNA, H/ACA box 74A (SNORA74A) reference gene were measured.
An enriched small RNA fraction was isolated using the mirVana miRNA isolation kit (Life Technologies, Auckland, NZ) according to the manufacturer’s instructions. For real-time PCR analysis, 10 ng of the small RNA fraction was reverse transcribed into cDNA using the TaqMan small RNA assay kit (Life Technologies, Auckland, NZ) according to the manufacturer’s instructions. The cDNA was stored at -20°C until required. The TaqMan assays used were; MIR107 (000443), MIR30E (002223), MIRLET7G (002282) and SNORA74A (001003). All PCRs were completed and analysed as described in the quantification of oxidative defence gene expression experiment.
Quantification of oxidative defence enzyme activity
The activity of the catalase, glutathione peroxidase, and superoxide dismutase oxidative defence enzymes in the RWPE-1 and WPE1-NA22 cell lines in response to 25 µM catechin or chlorogenic acid was measured using the absorbance-based CAT, SOD and GPx assay kits (707002, 706002 and 703102, Cayman Chemicals, Ann Arbor, MI, USA) according to the manufacturer’s instructions.
For each cell line, 5 x 105 cells were seeded into sufficient wells of a 6-well tissue-culture plate (Corning, Corning, NY, USA) flask and cultured for 24 hours to facilitate attachment. The medium was replaced with the appropriate supplemented (or control) media and cultured for 24 hours. CAT activity was measured at 540 nm, SOD activity was measured at 450 nm and GPx activity was measured at 340 nm (at 25°C every minute for 10 minutes) using a SpectraMax 250 spectrophotometer (Molecular Devices, Downington, PA, USA). The experiment was completed with six biological replicates each with thirty technical replicates for all samples.
Statistical analyses
All data were analysed for normality and statistical significance determined using ANOVA. Following analysis, significantly different means were identified using the least significant difference (LSD) post-hoc test. A P-value of less than 0.05 was considered to show a significant difference. All analyses were completed using GenStat v16 software (VSN International, Hertfordshire, UK).
Results
Free radical scavenging capacity assay
The effects of catechin, chlorogenic acid, and RWPE-1 and WPE1-NA22 cell lysates on free radical scavenging, as measured by DPPH radical formation, are shown in Figure 1. Catechin and chlorogenic acid significantly reduced the formation of the DPPH radical compared to the untreated sample (Figure 1a). Catechin was more effective than chlorogenic acid because it reduced DPPH radical formation at 1, 10, and 25 µM, whereas chlorogenic acid was only effective at 10 and 25 µM. Both the cell lysates are effective at reducing DPPH radical formation compared to the untreated sample (Figure 1b). When compared to the RWPE-1 cell lysate, only 10 and 25 µM catechin and 25 µM chlorogenic acid were more effective than the cell lysate alone Figure 1c). None of the chlorogenic acid concentrations tested were more effective that RWPE-1 or WPE1-NA22 cell lysates (Figure 1d).
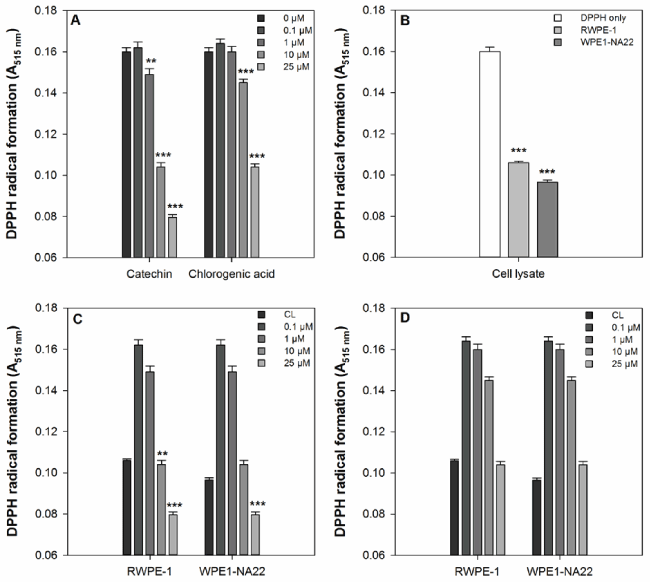
Figure 1. (a) The effect of catechin and chlorogenic acid compared to DPPH alone. A significant difference between treated and untreated samples is indicated by ** (P<0.01) or *** (P=0.001). (b) The effect of RWPE1- and WPE1-NA22 cell lysates compared to DPPH alone. A significant difference between treated and untreated samples is indicated by *** (P=0.001). (c) The effect of catechin on DPPH radical formation compared to RWPE-1 and WPE1-NA22 cell lysates. A significant difference between treated and untreated samples is indicated by *** (P=0.001). (d) The effect of chlorogenic acid on DPPH radical formation compared to RWPE-1 and WPE1-NA22 cell lysates. A significant difference between treated and untreated samples is indicated by *** (P=0.001).
Cell viability assay
Catechin or chlorogenic acid, at 25 µM, did not substantially or significantly alter the metabolic activity of RWPE-1 or WPE1-NA22 cells after 24 hours, indicating that this concentration of either compound does not alter the viability of these cell lines.
Cellular oxidative defence assay
The effect of 25 µM catechin or chlorogenic acid on the cellular oxidative defence of RWPE-1 and WPE1-NA22 cells, as measured by DCF formation, is shown in Figure 2. These data indicate that only catechin (at 25 µM), and not chlorogenic acid, reduces DCF formation in either cell line. There was no significant difference in the magnitude of catechin-mediated inhibition of DCF formation between the two cell lines.
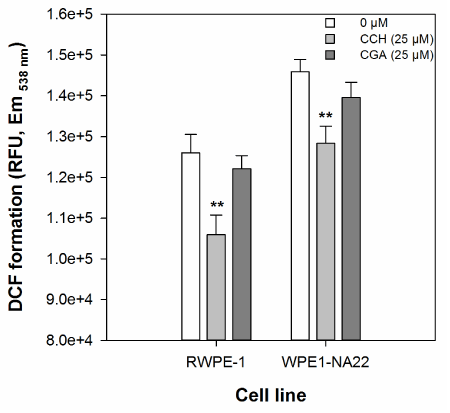
Figure 2. The effect of 25 µM catechin and chlorogenic acid on the cellular oxidative defence of RWPE1- and WPE1-NA22 cells, as measured by DCF formation. All data are expressed as the mean (±SEM) of twelve independent replicates. A significant difference between treated and untreated samples is indicated by ** (P<0.01).
The effect of catechin and chlorogenic acid on biological markers of the oxidative defence response and regulation by microRNA
The effect of 25 µM catechin or chlorogenic acid on the expression of the CAT, GPX1, and SOD2 anti-oxidant genes by RWPE-1 and WPE1-NA22 cells after 24 hours, as measured by probe-based real-time PCR, is shown in Figure 3a and 3d. These data indicate that CAT expression in the WPE1-NA22 cell line was affected by catechin or chlorogenic acid (reduced by 54 and 56%, respectively). Although not significant, the expression of the three genes was generally increased in the RWPE-1, and decreased in the WPE1-NA22 cell lines.
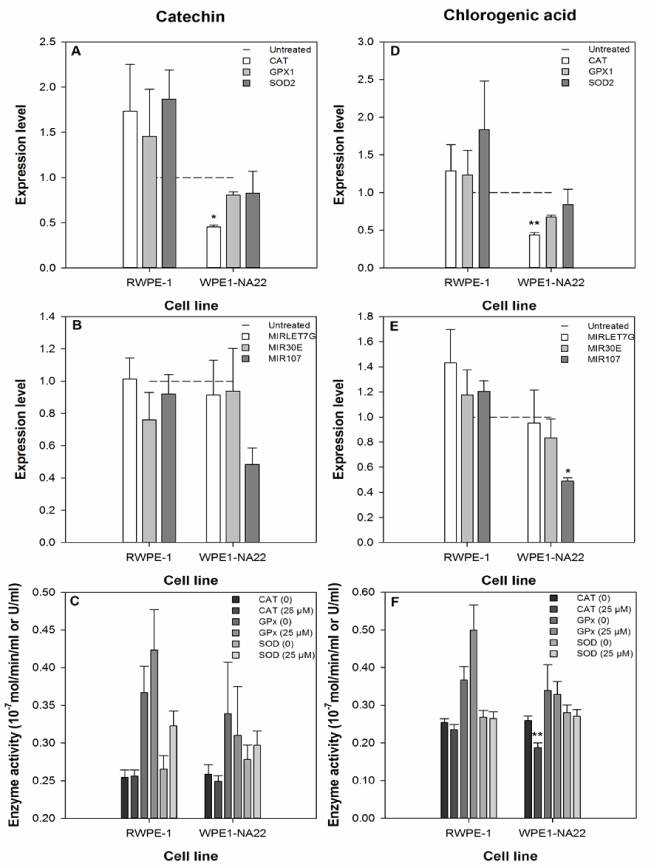
Figure 3. The effect of 25 µM catechin (a-c) and chlorogenic acid (d-f) on the expression and activity ofbiological markers of the oxidative defence response and expression of selected microRNAs. The expression level data are expressed as the mean (±SEM) of three independent replicates. A significant difference between treated and untreated samples is indicated by * (P<0.05). The enzyme activity data are expressed as the mean (±SEM) of six independent replicates. A significant difference between treated and untreated samples is indicated by ** (P<0.01).
The effect of 25 µM catechin or chlorogenic acid on the expression of the microRNAs MIRLET7G, MIR30E, and MIR107 (predicted to regulate the translation of the CAT, GPX1, and SOD2 anti-oxidant genes) by RWPE-1 and WPE1-NA22 cells after 24 hours, as measured by probe-based real-time PCR, is shown in Figure 3b and 3e. MIRLET7G is predicted to regulate CAT translation, MIR30E is predicted to regulate CAT and SOD2 translation, and MIR107 is predicted to regulate GPX1 and SOD2 translation. These data indicate that only the expression of MIR107 was affected by any treatment (reduced by 51% in chlorogenic acid treated WPE1-NA22 cells).
The effect of 25 µM catechin or chlorogenic acid on the activity of the CAT, GPx, and SOD enzymes in the RWPE-1 and WPE1-NA22 cell lines after 24 hours, is shown in Figure 3c and 3f. These data indicate that only the activity of the CAT enzyme was affected by any treatment (chlorogenic acid-treated WPE1-NA22 cells).
Discussion
Given the consumer interest in the phrase “anti-oxidant” [1,2] and that the primary method of assessing anti-oxidant activity is based on purely chemical assays [56], we developed a model as a tool to study how genetic, epigenetic, enzymatic, and phenotypic markers of the cellular response to oxidative damage may be altered by dietary factors. Catechin and chlorogenic acid were used as reference dietary factors because they are known anti-oxidants and can be readily consumed in our diet, e.g. coffee, tea, and chocolate. In order for our integrated method to be of value it is essential that it provides a more coherent and relevant interpretation of bio-activity than purely chemical screening alone.
The data show that although both catechin and chlorogenic acid act as anti-oxidants in a purely chemical in vitro assay (Figure 1); this is not the case in a biological in vitro anti-oxidant assay (Figure 2). These data indicate that using a cellular anti-oxidant assay provides more meaningful biological information than a chemical assay alone. This cellular anti-oxidant assay, developed by Wolfe et al. [57], is straightforward and can be completed in any standard cell culture facility. In our study we used the RWPE-1 cell line as a ‘normal’ cell line; however, it is not truly ‘normal’ because it is an immortalised cell line and is likely a mixture of cancer stem cells and non-cancer stem cells. The other cell line, WPE1-NA22, is derived from the RWPE-1 cell line and has a higher growth rate which is indicative of carcinogenic transformation [58,59]. We used these cell lines, not strictly for their carcinogenic state, but as relatively crude models of a healthy/less diseased and a more diseased cellular state. It is essential that the appropriate cell type, given the inherent limitations of cell culture, is used for screening nutrients for any bio-activity.
Our data show that chemical anti-oxidant activity is not necessarily predictive of biological activity and because of this we examined aspects of the cellular anti-oxidant systems to determine if measuring these added further value to the interpretation of biological response. At the transcriptional level, both catechin and chlorogenic acid substantially reduced the expression (~50%) of CAT in the WPE1-NA22 cell line (Figure 3a and d). The pattern of the response was similar for both cell lines, i.e. a tendency for increased expression in the RWPE-1 cell line and a decrease in the WPE1-NA22 cell line. This indicates that at the transcriptional level these cell lines appear not to differ sufficiently to function as appropriate models of ‘normal’ and diseased cellular states. Give the interest in the literature in the regulation of mRNA translation by microRNA, we assessed if the mRNA response we measured might be due to altered miRNA activity (Figure 3b and e). However, only the expression of MIR107 was affected by our treatments (chlorogenic acid reduced its expression in the WPE1-NA22 cell line). This is inconsistent with the mRNA data as MIR107 is predicted to target GPX1 and we measured no significant change in the expression of GPX1. Although microRNA may be involved in the regulation of the anti-oxidant response, we did not find sufficient evidence to justify its further inclusion in our integrated approach.
Logically, a change in mRNA abundance should lead to a change in protein abundance or activity so we measured the activity of the anti-oxidant enzymes encoded by the genes we studied (Figure 3c and f). We chose to measure activity rather than abundance because from a screening perspective, it is easier to do and it is the activity of the enzyme rather than just its abundance (although the two are clearly linked) that is more biologically relevant. We found that the activity of the catalase enzyme was reduced by chlorogenic acid in the WPE1-NA22 cell line. This is consistent with the reduction in CAT gene expression measured and indicates that when transcriptional data and enzyme activity data are considered together, it may provide a clearer interpretation of a biological response.
Although we have shown that chemical anti-oxidant activity does not necessarily predict biological activity, it is unclear from our data if measuring other biological activity, such as transcription, epigenetics, or enzyme activity substantially improves the interpretation of the cellular response to anti-oxidants. There is a disparity in our data in how the compounds tested respond biologically at the mechanistic level. Only catechin was an effective anti-oxidant in the cellular assay, but had no substantial effects on the mechanistic markers measured. Conversely, chlorogenic acid was ineffective in the cellular assay, but had some effects on the mechanistic markers measured.
However, the chemical and biological effects shown in Figures 1 to 3 occur at concentrations that may not be physiologically relevant. Both catechin and chlorogenic acid are extensively metabolised during digestion and absorption of foods containing them. The selection of a likely non-physiological concentration for study was based on the most effective concentration of both compounds in the chemical in vitro assay to maximise the assessment of our integrated approach.
These data show that despite both compounds having potent free radical scavenging capacity, this does not necessarily occur in biological systems. However, it is not clear how the amount of cellular uptake of these compounds or extracellular effects contribute to the reported effects, or absence of them. Several polyphenols have been shown to alter miRNA abundance [33,34,36-38,40-45,47,48,51,53] and the selection of dose in this study may not reflect physiological concentrations and exposure or the effect of metabolites. However, the primary aim of this study was to identify a concentration that was more effective chemically than cell lysates alone and to assess if using cellular and mechanistic tools improve the biological interpretation of anti-oxidant activity. We found that no single method (chemical, biological, or mechanistic) is satisfactory when assessing the bio-activity of nutrients. Furthermore, our data highlight the importance of selecting appropriate cell models and not relying on one screening method, chemical, cellular, or mechanistic, but rather an integrated approach consisting of at least a chemical and biological activity screening method.
Conclusions
In this study, a suite of tools was used to investigate the cellular aspects of the anti-oxidant activity of dietary ingredients, and provide exploratory mechanistic evidence for regulatory and marketing purposes. The effects of two well-characterised anti-oxidants, catechin and chlorogenic acid, were assessed in two cell lines, representing a model for normal and abnormal cell health. The application of assays to measure free radical scavenging and cellular oxidative defence enabled the anti-oxidant capacity of catechin and chlorogenic acid within the context of a biological system. Measurement of the expression and activity of key anti-oxidant genes, and miRNA molecules known to regulate these genes, were used to interrogate the potential mechanisms by which these compounds act as anti-oxidants.
These data show that although catechin and chlorogenic acid are potent anti-oxidants in a chemical assay, this activity does not necessarily occur in the cell models used. Although we found a limited role for epigenetic (miRNA) regulation of cellular response to oxidative stress, it cannot be excluded as a possible mechanism as other miRNAs and/or epigenetic factors may be involved. This study highlights the importance of using a range of techniques to properly characterise the effects of dietary ingredients.
Acknowledgments
The authors wish to acknowledge technical assistance from Ms. Ivy (Xuejing) Men.
Author contributions
M.M. completed data collection, sample analysis and data analysis. All authors contributed to the study design, review of the data, and preparation of the manuscript.
Conflicts of interest
2021 Copyright OAT. All rights reserv
The authors declare no conflict of interest. The authors alone are responsible for the content and writing of this article.
References
- EFSA Panel on Dietetic Products, N.a.A.N. (2011) Guidance on the scientific requirements for health claims related to antioxidants, oxidative damage and cardiovascular health. EFSA Journal 9: 13.
- EFSA Panel on Dietetic Products, N.a.A.N. (2010) Scientific opinion on the substantiation of health claims related to various food(s)/food constituent(s) and protection of cells from premature ageing, antioxidant activity, antioxidant content and antioxidant properties, protection of DNA, proteins and lipids from oxidative damage and bioavailability of anthocyanins in black currants pursuant to article 13(1) of regulation (ec) no 1924/2006. EFSA Journal 8: 34.
- Weng CJ, Yen GC (2012) Chemopreventive effects of dietary phytochemicals against cancer invasion and metastasis: phenolic acids, monophenol, polyphenol, and their derivatives. Cancer Treat Rev 38: 76-87. [Crossref]
- Williamson G, Dionisi F, Renouf M (2011) Flavanols from green tea and phenolic acids from coffee: critical quantitative evaluation of the pharmacokinetic data in humans after consumption of single doses of beverages. Mol Nutr Food Res 55: 864-873. [Crossref]
- Manach C, Williamson G, Morand C, Scalbert A, Rémésy C (2005) Bioavailability and bioefficacy of polyphenols in humans. I. Review of 97 bioavailability studies. Am J Clin Nutr 81: 230S-242S. [Crossref]
- Williamson G, Manach C (2005) Bioavailability and bioefficacy of polyphenols in humans. II. Review of 93 intervention studies. Am J Clin Nutr 81: 243S-255S. [Crossref]
- van der Burg-Koorevaar MC1, Miret S, Duchateau GS (2011) Effect of milk and brewing method on black tea catechin bioaccessibility. J Agric Food Chem 59: 7752-7758. [Crossref]
- Duarte GS, Farah A (2011) Effect of simultaneous consumption of milk and coffee on chlorogenic acids' bioavailability in humans. J Agric Food Chem 59: 7925-7931. [Crossref]
- Sato Y, Itagaki S, Kurokawa T, Ogura J, Kobayashi M, et al. (2011) In vitro and in vivo antioxidant properties of chlorogenic acid and caffeic acid. Int J Pharm 403: 136-138. [Crossref]
- Farah A, Monteiro M, Donangelo CM, Lafay S (2008) Chlorogenic acids from green coffee extract are highly bioavailable in humans. J Nutr 138: 2309-2315. [Crossref]
- Farrell TL, Dew TP, Poquet L, Hanson P, Williamson G (2011) Absorption and metabolism of chlorogenic acids in cultured gastric epithelial monolayers. Drug Metab Dispos 39: 2338-2346. [Crossref]
- Renouf M, Guy PA, Marmet C, Fraering AL, Longet K, et al. (2010) Measurement of caffeic and ferulic acid equivalents in plasma after coffee consumption: small intestine and colon are key sites for coffee metabolism. Mol Nutr Food Res 54: 760-766. [Crossref]
- Renouf M, Marmet C, Guy P, Fraering AL, Longet K, et al. (2010) Nondairy creamer, but not milk, delays the appearance of coffee phenolic acid equivalents in human plasma. J Nutr 140: 259-263. [Crossref]
- Stalmach A, Steiling H, Williamson G, Crozier A (2010) Bioavailability of chlorogenic acids following acute ingestion of coffee by humans with an ileostomy. Arch Biochem Biophys 501: 98-105. [Crossref]
- Katz DL, Doughty K, Ali A (2011) Cocoa and chocolate in human health and disease. Antioxid Redox Signal 15: 2779-2811. [Crossref]
- Pandey M, Gupta S (2009) Green tea and prostate cancer: from bench to clinic. Front Biosci (Elite Ed) 1: 13-25. [Crossref]
- Yang CS, Wang X, Lu G, Picinich SC (2009) Cancer prevention by tea: animal studies, molecular mechanisms and human relevance. Nat Rev Cancer 9: 429-439. [Crossref]
- Upham BL, Trosko JE (2009) Oxidative-dependent integration of signal transduction with intercellular gap junctional communication in the control of gene expression. Antioxid Redox Signal 11: 297-307. [Crossref]
- Valko M, Leibfritz D, Moncol J, Cronin MT, Mazur M, et al. (2007) Free radicals and antioxidants in normal physiological functions and human disease. Int J Biochem Cell Biol 39: 44-84. [Crossref]
- Hebert-Schuster M, Fabre EE, Nivet-Antoine V (2012) Catalase polymorphisms and metabolic diseases. Curr Opin Clin Nutr Metab Care 15: 397-402. [Crossref]
- Fukai T, Ushio-Fukai M (2011) Superoxide dismutases: role in redox signaling, vascular function, and diseases. Antioxid Redox Signal 15: 1583-1606. [Crossref]
- Miao L, St Clair DK (2009) Regulation of superoxide dismutase genes: implications in disease. Free Radic Biol Med 47: 344-356. [Crossref]
- Hempel N, Carrico PM, Melendez JA (2011) Manganese superoxide dismutase (Sod2) and redox-control of signaling events that drive metastasis. Anticancer Agents Med Chem 11: 191-201. [Crossref]
- Brigelius-Flohe R, Maiorino M (2012) Glutathione peroxidases. Biochim Biophys Acta.
- Lubos E, Loscalzo J, Handy DE (2011) Glutathione peroxidase-1 in health and disease: from molecular mechanisms to therapeutic opportunities. Antioxid Redox Signal 15: 1957-1997. [Crossref]
- Crawford A, Fassett RG, Geraghty DP, Kunde DA, Ball MJ, et al. (2012) Relationships between single nucleotide polymorphisms of antioxidant enzymes and disease. Gene 501: 89-103. [Crossref]
- Gülçin İ (2012) Antioxidant activity of food constituents: an overview. Arch Toxicol 86: 345-391. [Crossref]
- Richelle M, Tavazzi I, Offord E (2001) Comparison of the antioxidant activity of commonly consumed polyphenolic beverages (coffee, cocoa, and tea) prepared per cup serving. J Agric Food Chem 49: 3438-3442. [Crossref]
- Butt MS, Sultan MT (2011) Coffee and its consumption: benefits and risks. Crit Rev Food Sci Nutr 51: 363-373. [Crossref]
- Crozier TW, Stalmach A, Lean ME, Crozier A (2012) Espresso coffees, caffeine and chlorogenic acid intake: potential health implications. Food Funct 3: 30-33. [Crossref]
- Yazheng L, Kitts DD (2012) Activation of antioxidant response element (ARE)-dependent genes by roasted coffee extracts. Food Funct 3: 950-954. [Crossref]
- Boettler U, Sommerfeld K, Volz N, Pahlke G, Teller N, et al. (2011) Coffee constituents as modulators of Nrf2 nuclear translocation and ARE (EpRE)-dependent gene expression. J Nutr Biochem 22: 426-440. [Crossref]
- Shilo S, Roy S, Khanna S, Sen CK (2008) Evidence for the involvement of miRNA in redox regulated angiogenic response of human microvascular endothelial cells. Arterioscler Thromb Vasc Biol 28: 471-477. [Crossref]
- Xu Y, Fang F, Zhang J, Josson S, St Clair WH, et al. (2010) miR-17* suppresses tumorigenicity of prostate cancer by inhibiting mitochondrial antioxidant enzymes. PLoS One 5: e14356. [Crossref]
- Griffiths-Jones S (2006) miRBase: the microRNA sequence database. Methods Mol Biol 342: 129-138. [Crossref]
- Lançon A, Michaille JJ, Latruffe N (2013) Effects of dietary phytophenols on the expression of microRNAs involved in mammalian cell homeostasis. J Sci Food Agric 93: 3155-3164. [Crossref]
- Milenkovic D, Jude B, Morand C (2013) miRNA as molecular target of polyphenols underlying their biological effects. Free Radic Biol Med 64: 40-51. [Crossref]
- Chakrabarti M, Ai W, Banik NL, Ray SK (2013) Overexpression of miR-7-1 increases efficacy of green tea polyphenols for induction of apoptosis in human malignant neuroblastoma SH-SY5Y and SK-N-DZ cells. Neurochem Res 38: 420-432. [Crossref]
- Pan MH, Lai CS, Wu JC, Ho CT (2013) Epigenetic and disease targets by polyphenols. Curr Pharm Des 19: 6156-6185. [Crossref]
- Bladé C, Baselga-Escudero L, Salvadó MJ, Arola-Arnal A (2013) miRNAs, polyphenols, and chronic disease. Mol Nutr Food Res 57: 58-70. [Crossref]
- Izzotti A, Cartiglia C, Steele VE, De Flora S (2012) MicroRNAs as targets for dietary and pharmacological inhibitors of mutagenesis and carcinogenesis. Mutat Res 751: 287-303. [Crossref]
- Lançon A, Kaminski J, Tili E, Michaille JJ, Latruffe N (2012) Control of MicroRNA expression as a new way for resveratrol to deliver its beneficial effects. J Agric Food Chem 60: 8783-8789. [Crossref]
- Joven J, Espinel E, Rull A, Aragone, G, Rodriguez-Gallego E, et al. (2012) Plant-derived polyphenols regulate expression of mirna paralogs mir-103/107 and mir-122 and prevent diet-induced fatty liver disease in hyperlipidemic mice. Biochim Biophys Acta 1820: 894-899. [Crossref]
- Angel-Morales G, Noratto G, Mertens-Talcott S (2012) Red wine polyphenolics reduce the expression of inflammation markers in human colon-derived CCD-18Co myofibroblast cells: potential role of microRNA-126. Food Funct 3: 745-752. [Crossref]
- Milenkovic D, Deval C, Gouranton E, Landrier JF, Scalbert A, et al. (2012) Modulation of miRNA expression by dietary polyphenols in apoE deficient mice: a new mechanism of the action of polyphenols. PLoS One 7: e29837. [Crossref]
- Ho E, Beaver LM, Williams DE, Dashwood RH (2011) Dietary factors and epigenetic regulation for prostate cancer prevention. Adv Nutr 2: 497-510. [Crossref]
- Reuter S, Gupta SC, Park B, Goel A, Aggarwal BB (2011) Epigenetic changes induced by curcumin and other natural compounds. Genes Nutr 6: 93-108. [Crossref]
- McKay JA, Mathers JC (2011) Diet induced epigenetic changes and their implications for health. Acta Physiol (Oxf) 202: 103-118. [Crossref]
- Murase T, Misawa K, Minegishi Y, Aoki M, Ominami H, et al. (2011) Coffee polyphenols suppress diet-induced body fat accumulation by downregulating SREBP-1c and related molecules in C57BL/6J mice. Am J Physiol Endocrinol Metab 300: E122-133. [Crossref]
- Hoelzl C, Knasmüller S, Wagner KH, Elbling L, Huber W, et al. (2010) Instant coffee with high chlorogenic acid levels protects humans against oxidative damage of macromolecules. Mol Nutr Food Res 54: 1722-1733. [Crossref]
- Tsang WP, Kwok TT (2010) Epigallocatechin gallate up-regulation of miR-16 and induction of apoptosis in human cancer cells. J Nutr Biochem 21: 140-146. [Crossref]
- Lambert JD, Elias RJ (2010) The antioxidant and pro-oxidant activities of green tea polyphenols: a role in cancer prevention. Arch Biochem Biophys 501: 65-72. [Crossref]
- Link A, Balaguer F, Goel A (2010) Cancer chemoprevention by dietary polyphenols: promising role for epigenetics. Biochem Pharmacol 80: 1771-1792. [Crossref]
- Dudonne S, Vitrac X, Coutiere P, Woillez M, Merillon JM (2009) Comparative study of antioxidant properties and total phenolic content of 30 plant extracts of industrial interest using DPPH, ABTS, FRAP, SOD, and ORAC assays. J Agric Food Chem 57: 1768-1774. [Crossref]
- Huang D, Ou B, Prior RL (2005) The chemistry behind antioxidant capacity assays. J Agric Food Chem 53: 1841-1856. [Crossref]
- Moon JK, Shibamoto T (2009) Antioxidant assays for plant and food components. J Agric Food Chem 57: 1655-1666. [Crossref]
- Wolfe KL, Liu RH (2007) Cellular antioxidant activity (CAA) assay for assessing antioxidants, foods, and dietary supplements. J Agric Food Chem 55: 8896-8907. [Crossref]
- Webber MM, Quader ST, Kleinman HK, Bello-DeOcampo D, Storto PD, et al. (2001) Human cell lines as an in vitro/in vivo model for prostate carcinogenesis and progression. Prostate 47: 1-13. [Crossref]
- McCann MJ, Rowland IR, Roy NC (2013) Anti-proliferative effects of physiological concentrations of enterolactone in models of prostate tumourigenesis. Mol Nutr Food Res 57: 212-224. [Crossref]



